


Next:Printout
of estimated alleleUp:Interpreting
the text outputPrevious:Printout
ofContents
Printout of  when using prior population information
when using prior population information
When the program is run with prior population information, the printout
is a bit different. Here I have run the same data with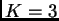 ,
USEPOPINFO=1, and GENSBACK = 1:
,
USEPOPINFO=1, and GENSBACK = 1:
Label (%Miss)
Pop :
1 17 (0)
2 : 0.998 | Pop 1: 0.000 0.001 | Pop 3: 0.000 0.000
2 1219 (7) 2 : 1.000
| Pop 1: 0.000 0.000 | Pop 3: 0.000 0.000
3 1223 (0) 2 : 0.612
| Pop 1: 0.000 0.000 | Pop 3: 0.004 0.383
4 1329 (7) 1 : 1.000
| Pop 2: 0.000 0.000 | Pop 3: 0.000 0.000
5 15 (0) 3 : 0.999
| Pop 1: 0.000 0.001 | Pop 2: 0.000 0.000
In general, the
first column of results (following ``Pop : '') shows the posterior probability
that the individual in question is correctly assigned to the given population.
The subsequent columns show the probabilities that it is from, or has ancestry
in, the other populations. There are GENSBACK+1 entries for each of the
other populations, showing the probability that an individual is from that
population, has a parent, grandparent, great-granparent,... etc, from that
population (in this order). For example, reading from row 3: Individual
1223 (who has 0% missing data) is actually from the presumed population
(2) with probability 0.612. There is (approximately) zero posterior probability
that this individual has recent ancestry in population 1, but it may have
recent ancestry in population 3 (the probabilities are 0.004, and 0.383,
that the individual is from population 3, or has a single parent
from population 3, respectively).



Next:Printout
of estimated alleleUp:Interpreting
the text outputPrevious:Printout
ofContents
William Wen 2002-07-18