



Next: How to cite this
Up: Interpreting the text output
Previous: Printout of estimated allele
Contents
When the SITEBYSITE option is chosen, the output file contains
posterior population of origin assignments for each allele copy at
each locus for each individual. For large datasets, this information
can require several megabases to store.
Each line shows the assignment probabilities for one locus for one
individual.The first two columns of the line indicate the number of
the individual (ranging from 1 to NUMINDS) and the number of the
locus (ranging from 1 to NUMLOCI) in the order that they occur in
the data file. If the data file contains labels for each individual or
marker names for each locus, these are given in subsequent
columns.
The format of the posterior assignment probabilities depends on the
parameter combinations. If LINKAGE=0 or PHASED=1 then the
first K rows of output give the probability that the first allele copy at
the locus comes from populations 1..K. For diploid or polyploid
data, analogous probabilities for subsequent allele copies are
shown in further columns.
If the linkage model is used (LINKAGE=1) and the data is not fully
phased (PHASED=0) the posterior assignment probabilites for the
allele copies at each locus can be strongly co-dependent.
Structure therefore outputs joint assignment probabilities for the
two allele copies implying 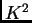 entries for each locus (note that this
option is not available for PLOIDY
entries for each locus (note that this
option is not available for PLOIDY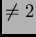 ).
If MARKOVPHASE=1 then the first K columns give the
probabilities that the first allele copy in the datafile is in population
1 and the second allele copy is in population 1..K, with subsequent
columns relating to probabilities with the first allele copy in
populations 2..K.
If MARKOVPHASE=0, then instead of referring to the first and
second listed allele copies in the data file, the probabilities refer to
the population of origin of maternal and paternal strands. If there is
no phase information (PHASEINFO=0), then the posterior
probability matrix should theoretically be symmetric, such that the
probability the maternal allele is in population
).
If MARKOVPHASE=1 then the first K columns give the
probabilities that the first allele copy in the datafile is in population
1 and the second allele copy is in population 1..K, with subsequent
columns relating to probabilities with the first allele copy in
populations 2..K.
If MARKOVPHASE=0, then instead of referring to the first and
second listed allele copies in the data file, the probabilities refer to
the population of origin of maternal and paternal strands. If there is
no phase information (PHASEINFO=0), then the posterior
probability matrix should theoretically be symmetric, such that the
probability the maternal allele is in population 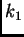 and the
paternal allele is in
and the
paternal allele is in 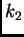 will be equal to the probability that the
maternal allele is in population
will be equal to the probability that the
maternal allele is in population 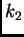 and the paternal allele is in
population
and the paternal allele is in
population 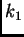 . In practice, because MCMC is used to estimate
the matrix, there will be noticeable deviations from symmetry if
NUMREPS is small.
. In practice, because MCMC is used to estimate
the matrix, there will be noticeable deviations from symmetry if
NUMREPS is small.




Next: How to cite this
Up: Interpreting the text output
Previous: Printout of estimated allele
Contents
William Wen
2002-07-18