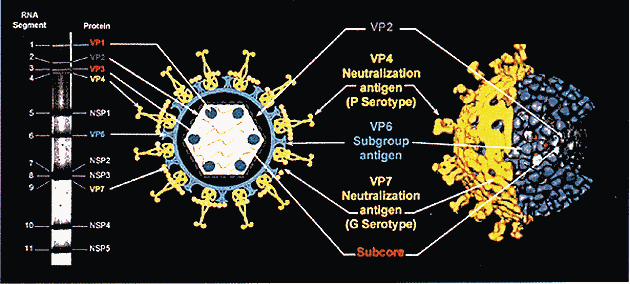
| Genome Segment |
Protein Producta |
Nascent Polypeptide (Mol. Wt× 1000)b |
Mature Protein Modified |
Location in Virus Particles |
Number of Molecules per virionc |
Ts mutant Groupd |
Functione |
| 1 |
VP1 |
125 |
- |
Core |
12 |
C |
RNA-dependent RNA polymerase, ss-RNA binding, complex with VP3 |
| 2 |
VP2 |
94 |
- |
Core |
120 |
F |
RNA binding, required for replicase activity of VP1 |
| 3 |
VP3 |
88 |
- |
Core |
12 |
B |
guanylytransferase, methytransferase, ss-RNA binding, complex with VP1 |
| 4 |
VP4 |
86.7 |
Cleaved VP5*(529) VP8*(247)f |
Outer capsid |
120 |
A |
Hemagglutinin, cell attachment, neutralization antigen, protease enhanced infectivity, virulence, putative fusion region, |
| 5 |
NSP1 (NS53) |
58.6 |
- |
Nonstructural |
|
NA |
Basic, zinc finger, RNA binding, virulence in mice; interacts with and degrades IRF-3; nonessential for some strains |
| 6 |
VP6 |
44.8 |
- |
Inner capsid |
780 |
G |
Hydrophobic, trimer, subgroup antigen, protection (intracellular neutralization ?); required for transcription |
| 7 |
NSP3 (NS34) |
34.6 |
- |
Nonstructural |
|
NA |
Acidic dimer, binds 3′ end of viral mRNAs, competes with cellular PABP for interaction with elF-4G1, inhibits host translation |
| 8 |
NSP2 (NS35) |
36.7 |
- |
Nonstructural |
|
E |
Basic, RNA binding, oligomer, NTPase, helicase, forms viroplasms with NSP5 |
| 9 |
VP7 |
37.4 |
cleaved signal peptide glycosylation |
Outer capsid |
780 |
NA |
RER intergral membrance glycoprotein, calcium-dependent trimer, neutralization antigen |
| 10 |
NSP4 (NS20) |
20.2 |
NS29–NS28 uncleaved signal peptide high mannose glycosylation and trimming |
Nonstructural |
|
NA |
RER transmembrance glycoprotein, intracellular receptor for DLPs, role in morphogenesis, interacts with viroplasms, modulates intracellular calcium and RNA replication, enterotoxin, secreted cleavage product, protection by antibody, virulence |
| 11 |
NSP5 (NS26) |
21.7 |
28K, O-glycosylated, phosphorylated (different forms) |
Nonstructural |
|
NA |
Basic phosphoprotein, RNA binding, protein kinase, forms viroplasms with NSP2, interacts with VP2 and NSP6 |
| |
NSP6 |
12 |
not known |
Nonstructural |
|
NA |
Interacts with NSP5, present in viroplasms and most virus strains |
aThe virion polypeptides are designated as proposed by Mason et al. (439) and modified by Liu et al. (418) and Mattion (449). VP3 is the protein product of gene segment 3, and VP4 is the protein product of gene segment 4. Early studies failed to unequivocally identify a protein product from genome segment 3; hence, the protein product of genome segment 4 was called VP3 in early publications. When the protein product of genome segment 3 was confirmed to be a structural protein located in the inner core, the genome 3 product was designated VP3 and the genome 4 product was renamed VP4 (418). In 1994, the nonstructural proteins were renamed NSP1 to NSP5 to facilitate comparisons among these proteins of different virus strains (449), and this nomenclature is now accepted. The parentheses in this table show the names of the nonstructural proteins as designated previously (NS followed by a number indicating their apparent molecular weight in thousands determined by electrophoresis in polyacrylamide gels containing sodium dodecyl sulfate). NSP6 has been characterized since 1994 (261,686).
bMolecular weights are for SA11 proteins and are calculated from the deduced amino acid sequences from the nucleotide sequence and from the longest potential open reading frame.
cCalculated from structural studies of purified virions (572,573,746).
dNA, no mutant assignment. Four groups of temperature sensitive mutants (tsD, tsH, I, J) are not yet mapped to a genome segment. Data compiled from the following sources: (111,259,437,582.)
eSee text for references.
fThree trypsin cleavage sites are found in SA11 VP4 at amino acids 231, 241, and 247. The indicated mature products are those based on use of only the preferred second cleavage site (420,422). |
|