


Next: Dominant loci
Up: Documentation for structure software:
Previous: Isolation by distance data
Contents
The program ignores missing genotype data when updating 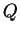 and
and 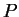 .
This approach is correct when the probability of having missing data
at a particular locus is independent of what allele the individual has
there. While estimates of
.
This approach is correct when the probability of having missing data
at a particular locus is independent of what allele the individual has
there. While estimates of 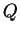 for individuals with missing data are
less accurate, there is no particular reason to exclude such
individuals from the analysis, unless they have very little data at
all.
A serious problem arises when data are missing in a systematic
manner, as with null alleles. These do not fit the assumed model, and
can lead to apparent departures from Hardy-Weinberg even without
population structure. One would not expect the assumed model to be
robust to this sort of violation.
Having multiple family members in the sample also violates the model
assumptions. Based on quite limited experience, my impression is that
this can sometimes lead to overestimation of
for individuals with missing data are
less accurate, there is no particular reason to exclude such
individuals from the analysis, unless they have very little data at
all.
A serious problem arises when data are missing in a systematic
manner, as with null alleles. These do not fit the assumed model, and
can lead to apparent departures from Hardy-Weinberg even without
population structure. One would not expect the assumed model to be
robust to this sort of violation.
Having multiple family members in the sample also violates the model
assumptions. Based on quite limited experience, my impression is that
this can sometimes lead to overestimation of 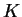 , but it may have
little effect on the assignment of individuals to populations for
fixed
, but it may have
little effect on the assignment of individuals to populations for
fixed 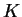 .
.
Subsections



Next: Dominant loci
Up: Documentation for structure software:
Previous: Isolation by distance data
Contents
Jonathan Pritchard
2003-07-10