
The replication scheme for the togavirus family is complex, yet fascinating, in that it served as a major distinguishing feature in its separate classification from the flavivirus family (they were once considered to be in the same family). Replication of the alphaviruses occurs in the cytoplasm, as in most RNA viruses, following uptake of the virion through receptor-mediated endocytosis:

The vesicles travel to the lysosomes where the lower pH induces release of the genome into the cytosol. Studies in-vitro andin-vivo have shown that binding of the capsid to ribosomes may also trigger the uncoating of the genome, allowing it to enter the cytoplasm and commence its translational activity. There also appears to be a ribosome-binding site on the capsid protein at the same spot where it interacts with the viral RNA during the encapsidation process. The genome is capped and polyadenylated at the 5' and 3' ends, respectively:
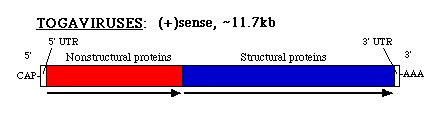
The 5' two thirds of the genome is translated into a large polyprotein (P1234)which cleaves itself and forms 4 nonstructural proteins (NSP1-methyalation and capping protein, NSP2-helicase and protease, NSP3-converts RNA replicaseto plus stranded replicase, NSP4-RNA polymerase). These proteins are indispensable for the production of a complementary RNA strand. NSP4 appears to be cleaved first and the P123/NSP4 complex is believed to initiate the synthesis of the complementary minus RNA strand. An interesting feature found in certain alphavirus species, such as Sindbis virus, is that a stop codon is found in front of the NSP4 sequence. A readthrough mechanism translates through this codon followed by cleavage of NSP4.
The complementary minus strand, formed by the NSP's, is used as a template for the production of a copy of the genomic RNA along with a subgeomic mRNA strand. The subgenomic message corresponds to the 3' end of the genome which contains the strucutural proteins (i.e. capsid proteins, evelope proteins, etc.) This allows for much greater efficiency in virion synthesis since one template strand can yield multiple copies of structural proteins, which are needed in great quantities to produce new virions. The subgenomic messge is then translated into another polyprotein which is also cleaved. Four or five structural proteins result from this cleavage: C-the nucleocapsid protein; E1,E2(some species also contain E3)-the envelope glycoproteins, and 6K-a small transmembrane protein.
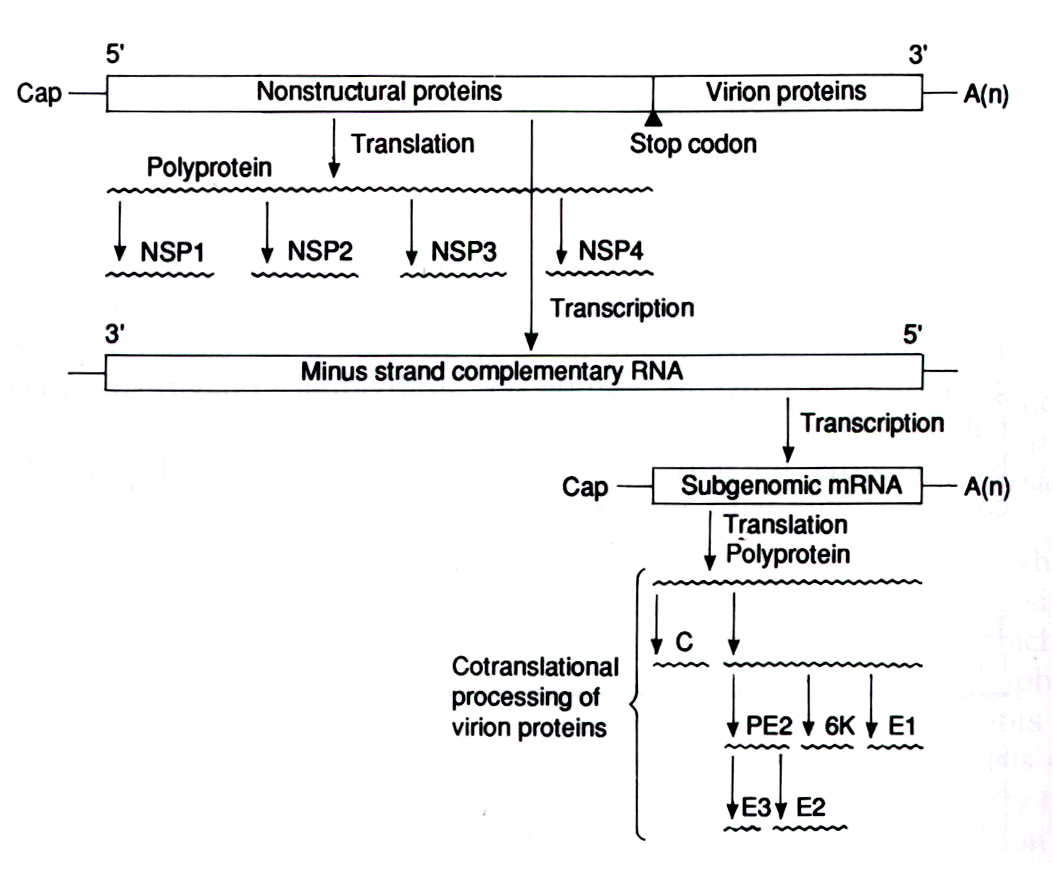
The capsid protein, C, is cleaved first through
the action of a serine-protease. The remaining polyprotein then migrates
into the lumen of the endoplasmic reticulum. Host cell signalases
cleave the 6K transmembrane protein along with the E1 protein from the
polyprotein. A complex known as P62(PE2), consisting of the
E2/E3
polyprotein, remains intact and travels to the plasma membrane
through secretory vesicles,
along with the E1 and 6K proteins. P62 will then be cleaved by a
furin-type protease which is located in trans Golgi vesicles.
Formation of new virions occurs in the
cytoplasm as genomic RNA and nucleoproteins assemble into a
nucleocapsid and then travel to the plasma membrane. Genomic RNA is
specifically selected to be surrounded by the capsid protein due to an
encapsidation signal, which is not found on subgenomic RNA.
The peplomer proteins found on the envelope are glycosylated as they exit
the endoplasmic reticulum, enter into the Golgi complex, and then
migrate to the plasma membrane. These proteins then attach to the
nucleocapsid and aid in the fusion of the capsid to the cell membrane,
thus leading to the budding of the newly enveloped virion. In
invertebrate cells the capsid buds through intracytoplasmic membranes and
is then expelled into the extracellular fluid.
Replication of the rubiviruses (rubella) is similar to that of alphaviruses in that they also produce a subgenomic mRNA message which codes for structural proteins at the 3' end of the genome, while the 5' end codes for nonstructural proteins. They do differ, however, in their strategy for cleaving the subgenomic polyprotein as well as in assembly of new virions. The capsid protein is not self-cleaved in the rubivirus genus, but is instead cleaved by a host cell signalase. The capsid protein is first cleaved from the E1 protein and remains attached to E2. While in the endoplasmic reticulum, C and E2 are cleaved and the capsid protein attaches to the membrane. Migration of the envelope glycoproteins from the ER to the Golgi is extremely slow. The majority of these proteins remain in the Golgi and very few reach the plasma membrane. This may be an indication as to why viral assembly predominantly occurs here (in the Golgi).