About Our Lab
Upcoming for 2026
Lab Update 2025
Lab Update 2024
Lab Research 2019 through 2016
Lab Research 2015 through 2013
Lab research 2012 and earlier
Our NSF-funded project
Early Anthers
Cell Fate Acquisition in Anthers
Why corn?
Detecting Mutator activity
Transposon tagging and gene cloning
Corn field at San Luis Obispo
UVB and Mu elements
Ustilago maydis
Male-sterile mutants
Learn more:
Transcriptome profiling in pre-meiotic and tapetal stages
Transcriptome profiling across all anther stages
Small RNAs in rice and maize anthers
Mu activity reprograms maize anther proteome and transcriptome
ms8 causes precocious anther development
Impact of Ustilago maydis on maize flowers
Organ specificity in Ustilago-maize interaction
Emergence and patterning of cell types in maize anthers
Learn more:
Transcriptome profiling in pre-meiotic and tapetal stages
Transcriptome profiling across all anther stages
Small RNAs in rice and maize anthers
Mu activity reprograms maize anther proteome and transcriptome
ms8 causes precocious anther development
Impact of Ustilago maydis on maize flowers
Organ specificity in Ustilago-maize interaction
Emergence and patterning of cell types in maize anthers
Technologies utilized:
2D DIGE
PALM (Positional Ablation w/ Laser Microbeams)
Oligo microarrays
Confocal microscopy

NEWS FLASH: Virgina Walbot awarded the 2023 Barbara McClintock Prize in Genetics and Genomics.
High Level Overview
We are studying 90 hours of early anther development in maize (Zea mays L., “corn”) to elucidate the signaling networks that regulate cell type acquisition and maintenance within anther locules. The question: Without a germ line, how do cells in plants switch from mitosis to meiosis?What's new in 2023?
Single cell RNA-seq with plantsImplementing single cell RNA-seq (using the Cell Seq2 protocol and new methods for isolating plant cells, see Marchant et al. doi.org/10.1101/2021.10.11.463960.) has fueled recent discoveries and publications. In 2019 Nelms & Walbot 10.1126/science.aav6428 including the cover picture traced germinal cell ontogeny in maize anthers, from the mitotic proliferating archesporial cells, through meiotic commitment to the leptotene/zygotene transition in prophaseI. This landmark study was one of the very first successes of single cell RNA-seq with plant materials. In this article, we demonstrated the method reproducibility by splitting the RNA extracted from each germinal cell to generate paired technical replicates - a first in any organism. We also implemented a new analytical method - pseudotime velocity - the pinpoint stages of rapid transcriptome reorganization, both gain and loss of transcript types. The analysis of 144 carefully staged cells established that all archesporial cells proliferate (rather than their being just a few proliferative stem cells) and that there is a modest transcriptome change as archesporial cells differentiate as Pollen Mother Cells, and then a dramatic transcriptome change at the L/Z boundary. In 2022 we defined 10.1126/science.abl7392 (again with a cover photo!) subsequent stages of germinal development through completion of meiosis, uninucleate microspore biogenesis, binucleation, and trinucleate pollen maturation.

Haploid gametophyte gene expression, a key feature of the alternation of generations, is hypothesized to reduce genetic load by selection against deleterious mutations for gene expression shared with the diploid sporophyte. This article enumerates the substantial genome fraction subject to this purifying selection. The shocking surprise, however, was that stored (carryover) mRNAs synthesized during diploidy, likely early in meiosis, persist as endowment in the haploid uninucleate microspores without contributions from the haploid genome for 11 days post-meiosis. Haploid gene expression is activated during the transition to the binucleate microspore.
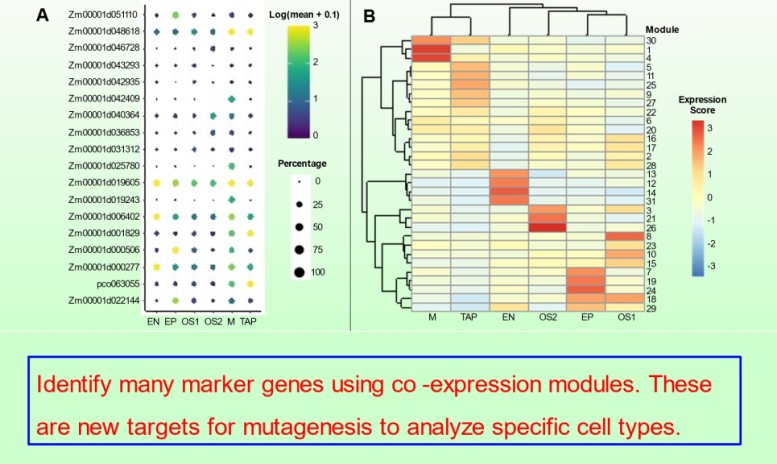
bHLH Transcription factors and development of the tapetum
We continued our studies of bHLH transcription factors that modulate tapetal development in maize anthers in Nan et al. 10.1093/plcell/koac007. The MS23, MS32, and bHLH122 proteins regulate 24-nt phasiRNA production in the tapetum in addition to sequentially coordinately tapetal redifferentiation steps prior to and during meiosis.
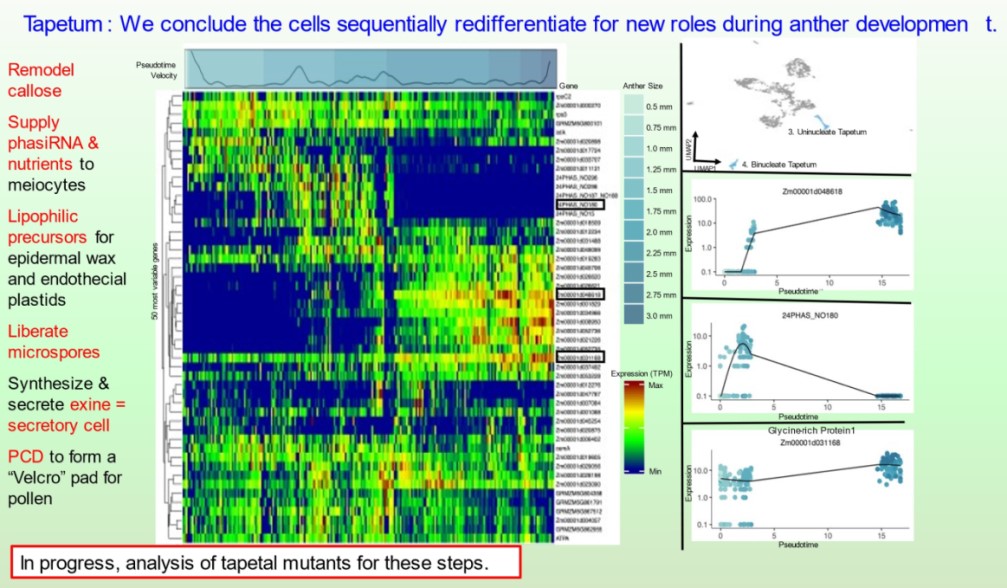
Subsequently, the bHLH51 factor regulates tapetal support of uninucleate microspore development. To further understand the tapetum and other somatic cell types, postdoc Blaine Marchant has completed single cell RNA-seq starting prior to the birth of tapetal and middle layer cells through the start of tapetal programmed cell death 3 weeks later. We discuss our views of anther development in 10.1093/plcell/koac287 a review prepared for a special issue celebrating Gregor Mendel. Implementing single cell RNA-seq of maize leaf protoplasts transiently expressing anther transcription factors is a new approach to organizing regulatory hierarchies for maize somatic anther cells 10.1111/tpj.16045.
2 new studies on 24-nt phasiRNA
Two published studies established new aspects of 24-nt phasiRNA biology. Zhang et al. 10.1111/nph.17060 demonstrated that these small RNAs are required for increased CHH DNA methylation at their own loci. This was readily detected in whole anthers at a specific stage of anther development, meaning that many somatic cells experience this. As the 24-nt phasiRNAs are produced in the tapetum, we hypothesize that tapetal cell DNA is impacted. This extends a previous literature report that the rare meiocyte cells show increased CHH DNA methylation at the 24-PHAS loci. Using laser microdissection to recover purified germinal, tapetal, and other somatic cells, Zhou et al. 10.1111/nph.18167 confirmed that 24-PHAS loci are expressed primarily in the tapetum, the 24-nt phasiRNAs are generated there, but by late meiosis I, these small RNAs have accumulated to high levels in the meiocytes. This fits nicely with the previous observation that meiocyte 24-PHAS loci experience hyper CHH DNA methylation. In on-going work, we are attempting to determine which, if any, Argonaute proteins are required for the action of 24-nt phasiRNAs, and of course, we are actively trying new experimental approaches to defining how 24-nt phasiRNAs modulate anther development.
Temperature-sensitive male fertility and dcl5
Although temperature-sensitive male fertility has been widely reported in a number of crop plants, it was not a critical feature in maize, except under extreme heat or drought conditions in which the tassel is "fried" and killed. Nonetheless, we discovered that the deficiency in dcl5, the gene required for processing the 24-PHAS precursor transcripts into 24-nt phased small RNAs, confers temperature-sensitive fertility. Under standard maize growing conditions, plants are full male-sterile, however, slow growth in cool conditions permits fertility restoration 10.1038/s41467-020-16634-6. Surprisingly, the well-studied transcription factor OCL4 conditions temperature sensitivity: in this case high growth temperature conditions some fertility restoration 10.1007/s00497-021-00406-3 Pursuing how temperature influences specific molecular and biochemical steps in anther development is a goal of our on-going work. Our working hypothesis is that 24-nt phasiRNAs participate by buffering anther development against unfavorable environmental conditions. Defining how this is accomplished is a major challenge.
Other items
We also published a brief review on maize - Ustilago interaction 10.1007/s00497-021-00406-3 and as part of a large consortium, an assessment of a teaching tool in which students learn gene annotation in a contest pitting one school against another. 10.1371/journal.pcbi.1009463 Stanford won it's rounds and inspired several students to pursue further training in bioinformatics!!