


Next:OverviewUp:IntroductionPrevious:Registering.Contents
What's new in Version 2?
Perhaps the most obvious addition to structure in Version 2 is a
fancy Java front end. The front end includes several new features, producing
various plots of data summaries, as well as enabling batch runs, and organizing
the data from multiple runs on the same project. C executables will still
be available for those who prefer using the command line. See section 7.
We have added command line options to simplify simulation studies and batch
runs (section 6.6). On the
analysis side, the major advance is that we have now implemented a model
that allows for ``admixture linkage disequilibrium''. That is, we model
the correlations that arise between linked markers as the result of admixture
or hybridization. In order to use this, the user must enter map distances
(or at least some kind of relative distance) between the markers, and the
program then accounts for the possibility of correlations due to linkage.
We still do not attempt to model the LD that occurs within populations
between very nearby markers (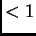 cM in humans). See section 3.2.
We have also modified and improved the model of correlated allele frequencies,
and we find that this model is often more effective than the independent
frequencies model at detecting subtle population structure. However, this
may come at some cost: we believe that it may be less robust to minor model
departures like some inbreeding, or undetected null alleles. This could
create a bias towards overestimating the number of clusters,
cM in humans). See section 3.2.
We have also modified and improved the model of correlated allele frequencies,
and we find that this model is often more effective than the independent
frequencies model at detecting subtle population structure. However, this
may come at some cost: we believe that it may be less robust to minor model
departures like some inbreeding, or undetected null alleles. This could
create a bias towards overestimating the number of clusters, 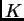 .
We have made the new model the default model, but encourage users to experiment
with different models. Other new extensions include the possibility of
using different
.
We have made the new model the default model, but encourage users to experiment
with different models. Other new extensions include the possibility of
using different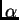 s
for each population, and estimating
s
for each population, and estimating 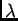 s.
The computational techniques underlying the new analytical methods are
described in a forthcoming paper (Falush
et al ., 2003) We have also modified the program so that it can be
applied to non-diploid organisms. As Daniel Falush points out, this should
keep the dandelion population geneticists happy.
s.
The computational techniques underlying the new analytical methods are
described in a forthcoming paper (Falush
et al ., 2003) We have also modified the program so that it can be
applied to non-diploid organisms. As Daniel Falush points out, this should
keep the dandelion population geneticists happy.



Next:OverviewUp:IntroductionPrevious:Registering.Contents
William Wen 2002-07-18