


Next:Columns:Up:Format
for the dataPrevious:Components
of the data
Contents
Row
-
Marker Names (Optional; string) The first row in the file can contain
a list of identifiers for each of the markers in the data set. This row
contains
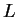 strings of integers or characters, where
strings of integers or characters, where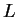 is the number of loci.
is the number of loci.
-
Inter-Marker Distances (Optional; real) the next row in the file
is a set of inter-marker distances, for use with linked loci. These should
be genetic distances (e.g., centiMorgans), or some proxy for this based,
for example, on physical distances. The actual units of distance do not
matter too much, provided that the marker distances are (roughly) proportional
to recombination rate (the algorithm estimates an appropriate scaling from
the data). The markers must be in map order within linkage groups. When
consecutive markers are from different linkage groups (e.g., different
chromosomes), this should be indicated by the value -1. The first marker
is also assigned the value -1. All other distances are non-negative. This
row contains
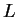 real numbers.
real numbers.
-
Individual Data (Required) Data for each sampled individual is arranged
into one or more rows as described above (further details below).
-
Phase Information (Optional; diploid data only; real number in the
range [0,1]). This is for use with linked loci only. This is a single row
of
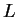 probabilities that appears after the genotype data for each individual.
If phase is known completely, or no phase information is available, these
rows are unnecessary. They may be useful when there is partial phase information
from family data or when haploid X chromosome data from males and diploid
autosomal data are input together. There are two alternative representations
for the phase information: (1) the two rows of data for an individual are
assumed to correspond to the paternal and maternal contributions, respectively.
The phase line indicates the probability that the ordering is correct at
the current marker (set MARKOVPHASE=0); (2) the phase line indicates the
probability that the phase of one allele relative to the previous allele
is correct (set MARKOVPHASE=1). The first entry should be filled in with
0.5 to fill out the line to
probabilities that appears after the genotype data for each individual.
If phase is known completely, or no phase information is available, these
rows are unnecessary. They may be useful when there is partial phase information
from family data or when haploid X chromosome data from males and diploid
autosomal data are input together. There are two alternative representations
for the phase information: (1) the two rows of data for an individual are
assumed to correspond to the paternal and maternal contributions, respectively.
The phase line indicates the probability that the ordering is correct at
the current marker (set MARKOVPHASE=0); (2) the phase line indicates the
probability that the phase of one allele relative to the previous allele
is correct (set MARKOVPHASE=1). The first entry should be filled in with
0.5 to fill out the line to 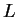 entries.For example the following data input would represent the information
from an male with 5 unphased autosomal microsatellite loci followed by
three X chromosome loci, using the maternal/paternal phase model:
entries.For example the following data input would represent the information
from an male with 5 unphased autosomal microsatellite loci followed by
three X chromosome loci, using the maternal/paternal phase model:
| 102 |
156 |
165 |
101 |
143 |
105 |
104 |
101 |
|
|
| 100 |
148 |
163 |
101 |
143 |
-9 |
-9 |
-9 |
|
|
| 0.5 |
0.5 |
0.5 |
0.5 |
0.5 |
1.0 |
1.0 |
1.0 |
|
|
| |
|
|
|
|
|
|
|
|
|
where -9 indicates "missing data", here missing due to the absence of a
second X chromosome, the 0.5 indicates that the autosomal loci are unphased,
and the 1.0s indicate that the X chromosome loci are have been maternally
inherited with probability 1.0, and hence are phased. The same information
can be represented with the markovphase model.In this case the input file
would read:
| 102 |
156 |
165 |
101 |
143 |
105 |
104 |
101 |
|
|
| 100 |
148 |
163 |
101 |
143 |
-9 |
-9 |
-9 |
|
|
| 0.5 |
0.5 |
0.5 |
0.5 |
0.5 |
0.5 |
1.0 |
1.0 |
|
|
| |
|
|
|
|
|
|
|
|
|
Here, the two 1.0s indicate that the first and second, and second and third
X chromosome loci are perfectly in phase with each other. Note that the
site by site output under these two models will be different. In the first
case, structure would output the assignment probabilities for maternal
and paternal chromosomes. In the second case, it would output the probabilities
for each allele listed in the input file.



Next:Columns:Up:Format
for the dataPrevious:Components
of the dataContents
William Wen 2004-07-13