Introduction to Scalable Neuroscience: Part 1
 |
Welcome to Stanford CS379C. If you haven't yet seen the poster or read the syllabus, please take a moment to read it here where you'll also find some information that might prove useful in case you're thinking about taking the course for credit. I'll be making presentations and leading discussions in the first two classes. With the exception of a few classes relating to projects, the remaining classes will feature scientists and engineers whose work is directly relevant to the challenge issued in the syllabus. You can check out the list of participants on the class calendar here.
This first presentation and class discussion focus on some of the current challenges in molecular and cellular neuroscience, emphasizing factors that influence the neuroscience projects that I work on at Google. We'll discuss what makes the most sense in terms of developing new technologies that accelerate the science, and talk about why this might be a good time to deviate from the trend toward incremental science and technology development and take on some really challenging problems that have the potential to completely change the next generation of digital prostheses and personal assistants.
The second presentation and class discussion introduce the programmer's apprentice project, the role it played in the early history of AI and the role it will play in this class, as well as some of the reasons why this is a perfect time to consider such an ambitious project. I'll provide a preview of several core technologies that we'll be looking at in class and introduce you to some of the relevant research in cognitive and systems neuroscience, machine learning and artificial neural networks, much of it originating from the technology companies and research laboratories of our invited speakers.
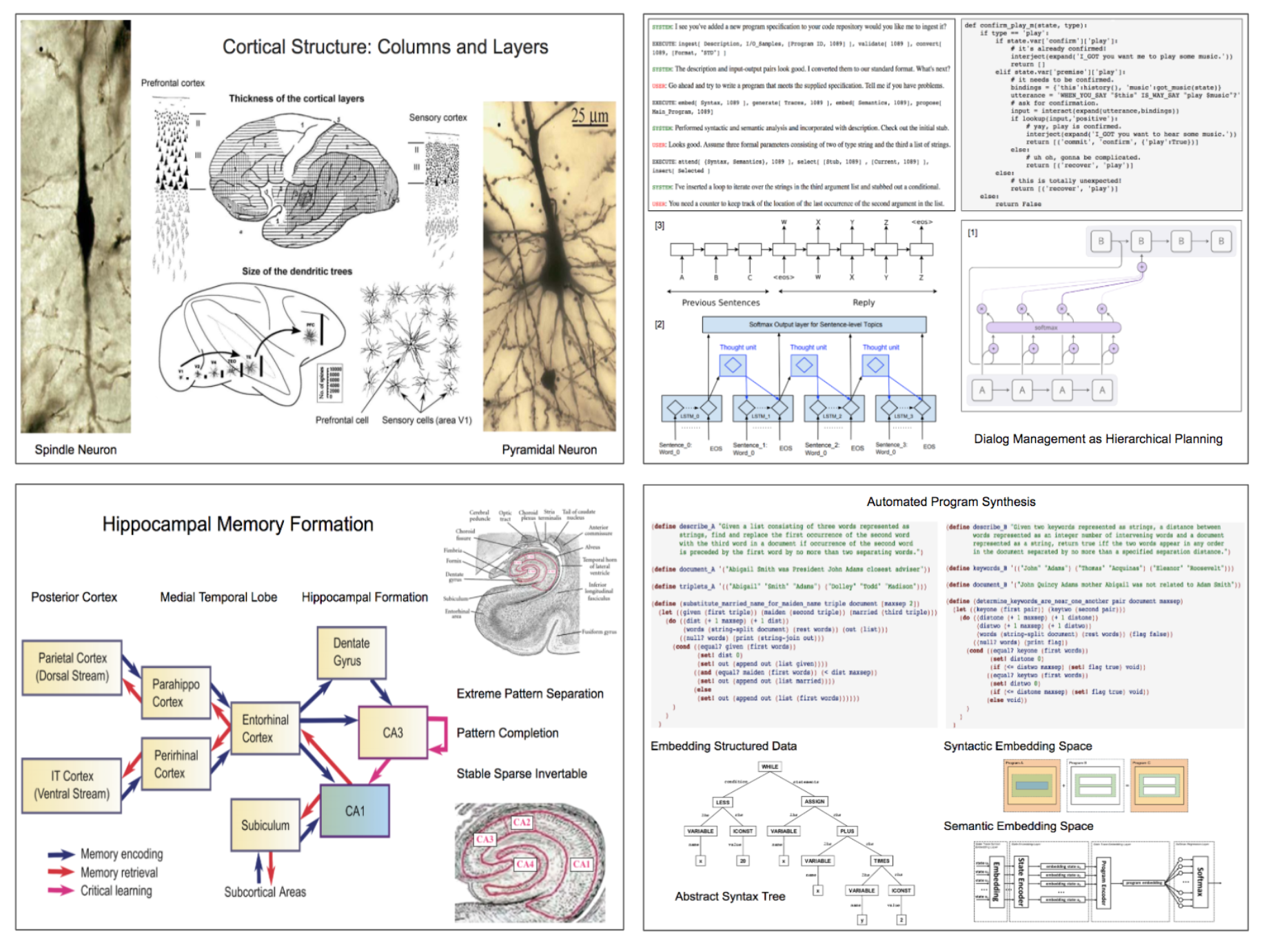 |
The four panels shown here demonstrate how applications drive the search for cognitive strategies and how concepts from cognitive neuroscience provide ideas for designing neural network architectures. The two panels on the right illustrate input-output modalities important for the programmer's apprentice. While a day will come when we can directly interface biological brains and digital prostheses, natural language offers the current best interface between the programmer and apprentice. We'll recast dialog management as hierarchical planning and demonstrate how it can be implemented using ideas from machine translation.
We are making substantial progress using differential models for automated program synthesis. The panel on the lower right illustrates how we might support a form of programming by code search substitution using sequence-to-sequence embedding models. At the core of this approach is the method of embedding both syntactic and semantic encodings of programs. Machine translation technologies can be used to embed the abstract syntax tree (AST) representations of programs. Multi-stage embeddings can be used encode semantic representations produced from code traces generated by running programs on sample input examples.
The two panels on the left illustrate issues relating to memory that arise in the programmer's apprentice. The top left is one of several slides describing Stanislas Dehaene's global workspace theory explaining how complex representations are constructed in short term memory using attentional mechanisms that can be implemented using neural network architectures. The bottom left is relevant to the problem of having new memories corrupt older memories, in particular those corresponding to episodic memory that record our thoughts and activity, especially in the context of our social interactions. Each panel represents a potential project.
 |
The human brain is very complicated. Some scientists claim that it is the most complicated artifact in the universe, but I doubt we've seen enough of the universe to say this with any confidence. At any rate, we have a long way to go in order to understand the human brain all the way down to the cellular and molecular levels. Understanding our biological brains is not just important from a scientific perspective.
Most of us know someone who is suffering from some form of brain disorder: a sibling with autism or attention deficit disorder, a parent with symptoms of early onset dementia or a grandparent suffering the ravages of late-stage Alzheimer's disease. You or one of your friends may be afflicted with mood disorders. These are devastating diseases that we would all like to find cures for as quickly as possible.
I believe that practical human intelligence by which I mean the ability solve complex scientific and engineering problems is not nearly as complicated as understanding the brain. Decades of research in cognitive and systems neuroscience have provided us with extraordinary insights into how human beings go about solving problems — insights some of us believe will enable us to replicate and then improve on human problem solving.
I believe we can replicate practical human intelligence and improve upon nature not just in terms of speed and accuracy, but also in terms of technical and scientific innovation. This is not the same thing as saying we can replicate human cognition or that we can build a robot companion that can understand another human being as well as a human companion. These are interesting topics to think about, but our goal here is not to replicate or replace humans.
Our grand goal is to build intelligent prostheses to help us understand our brains in order to eradicate suffering and give every human a chance to realize their potential. It isn't that building intelligent systems is better than studying neuroscience or medicine. The rationale is simply that it is more efficient for some of us to build better thinking machines so that we can accelerate our primary goal of understanding ourselves.
Many people believe that human beings are the most intelligent inhabitants on this planet. You may be the last generation to be able to reasonably entertain such a conceit. Perhaps with the advent of machines that think significantly better and faster than humans who score the highest on intelligence tests, win math Olympiads or receive Nobel Prizes, we will finally learn to appreciate admirable human traits that are even more rare in the universe than intelligence.
 |
I'm going to start with a brief reprise of a talk I gave at the Kavli Foundation Future Symposium on Technology to Accelerate Research in Neuroscience which was held in Santa Monica last October. The talk was intended to be controversial in order to spur discussion of several critical issues facing the field. I attempted to channel the perspective of industrial program managers and federal funding agencies in asking hard questions about how best to to allocate scarce resources.
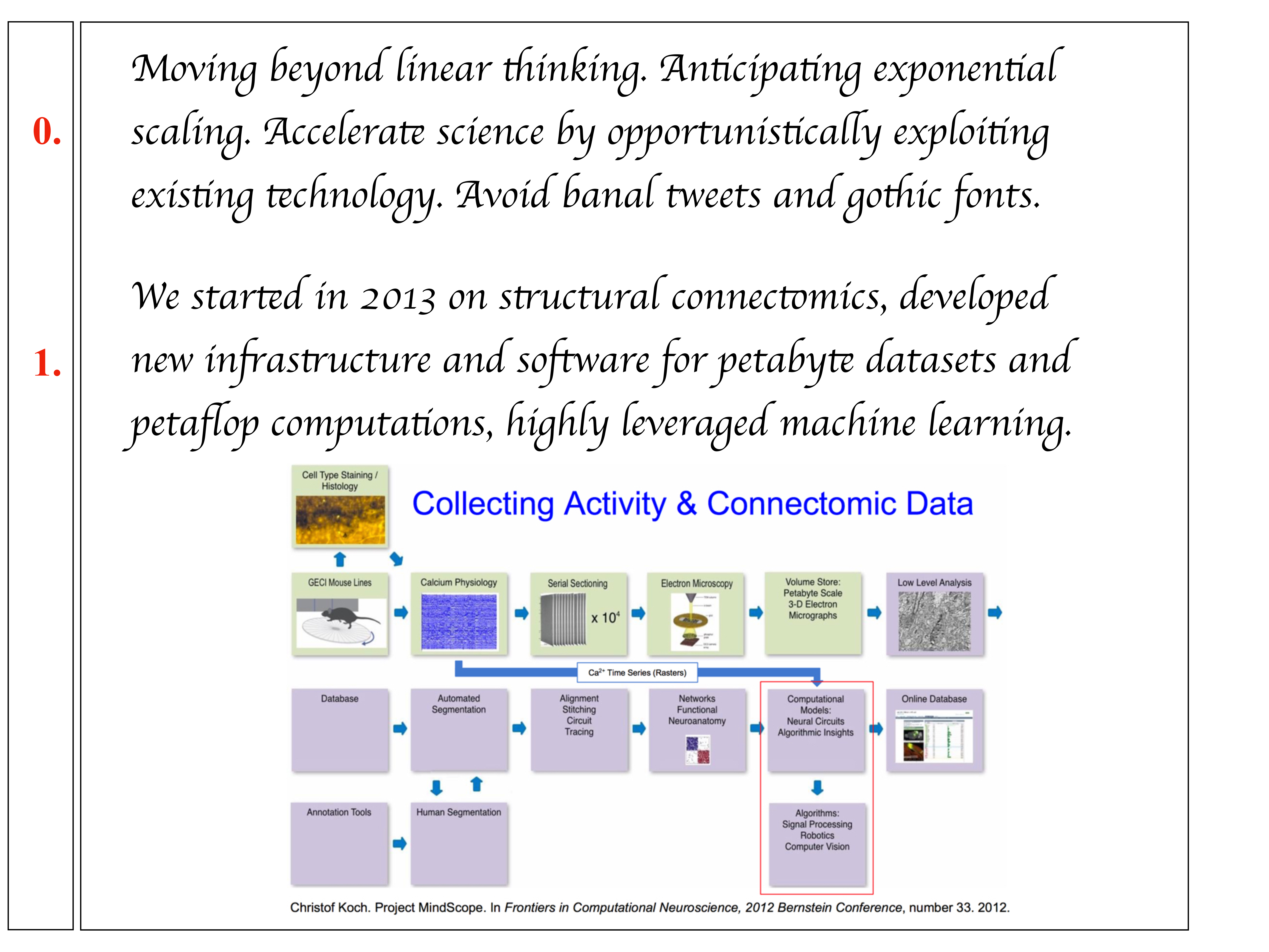
Human beings are used to a world in which almost everything scales linearly. This mindset is built deeply into the human psyche and has served us well throughout our relatively short evolutionary history. However, we are now entering a phase in which humans are learning how to exploit both natural processes and synthetic technologies in order to achieve exponential returns. This trend is relevant to our discussion today because the technologies we will be talking about in this class will accelerate that trend.
In 2013 we started work developing technology that would turn electron micrographs of neural tissue into complete wiring diagrams starting with the mouse visual cortex. This work was in collaboration with the Allen Institute for Brain Science and in particular with Christof Koch and Clay Reid. Generating these wiring diagrams — which are called connectomes — was but one part of a larger challenge that also required using two-photon microscopy1 to record the activity of every neuron within the target neural tissue in an awake behaving organism. The grand challenge was to understand the mouse visual system.
 |
In the version of this class taught in the Spring of 2013, the students and I wrote a joint paper entitled Technology Prospects for Scalable Neuroscience in which we concluded — along with other milestones — that within 2 to 5 years we would be able to generate the complete connectome for an entire organism. It has taken somewhat longer than we expected, but we were optimistic and there were quite a few Rumsfeld unknowns — things we didn't know that we didn't know — in particular, we didn't account for the many ways in which technology projects can fall prey to the vicissitudes of both human bureaucracies and the unrelenting laws of basic physics.
 |
This animation shows a simulated flight through a reconstructed neural tissue sample. It starts out looking at grayscale images corresponding to individual micrographs 100,000 pixels on a side. As we zoom in you can see the length of the scale bar start at around 10 microns and quickly shrinking to less than one micron. Next we view the segmentation of an individual micrograph in which different colors are assigned to each neurite to distinguish individual cells.
The segmented patches correspond to neurons, glia and blood vessels. The resolution is on the order of 10 nm in the X and Y plane of the micrograph. The tissue sample is sliced into sections between 20 and 40 nm thick thereby establishing the resolution in the Z plane. Larger tissue samples on the order of 50-100TB are cut up into smaller volumes 100 μm on a side. These volumes are imaged separately and then aligned and spliced back together prior to segmentation and tracing.
The animation shows fully reconstructed traces emerging out of the sample with yellow markers indicating the location of synapses. The graphics don't begin to do justice to the level of detail available. Only a very small number of neurons are actually displayed since if we were to display them all there would be little to see except the parts of processes that protrude outside of the tissue block. In addition to the reconstructed membranes of the neurons, we can resolve many of the organelles within individual cells including nuclei and mitochondria and even estimate the number of vesicles in the vicinity of individual synapses.
You may have seen similar animations in the past. What sets this apart is that it is a full reconstruction of a tissue sample significantly larger and more accurately traced than ever achieved prior. Moreover, the pipeline that produced it is fully automated, even though at this stage in its development we still employ human proofreaders to perform spot checks since neural tissue varies significantly across different organisms and we have much to learn about this variability. While we provide much of the core technology, the overall effort requires the collaboration of several labs and dozens of scientists.
The technology that generated the reconstruction consists of much more than machine learning and computer vision algorithms. We depend enormously on our colleagues at Google and at institutions around the world for new ideas about how to build systems of the sort we develop. We also invented and made open-source additional tools and technologies, but we would never have achieved what we have as quickly as we have were it not for the huge data centers and powerful computing infrastructure that Google researchers and product teams rely on to deliver new products and applications.
 |
At the symposium I asked the question, given what we know now, if we didn't have the search engines powered by data centers leveraging millions of cores, high-speed networking and exabytes of fast storage, how would we best spend our time and energy if our goal was to understand biological computation and the human mind. With 20:20 hindsight, my answer is that we would invent the World Wide Web, index all of the world's knowledge and build powerful search engines and the data centers necessary to run them, to accelerate the pace at which we learn about and understand the world around us.
 |
In 2015, I started work on developing technology for learning mesoscale models of neural circuits from a combination of structural and functional data. I was confident even then that we could obtain the necessary structural information, and I had started collaborations with scientists all over the world to better understand the prospects for obtaining the necessary whole-brain, pan-neuronal functional activity recordings.
The idea of spending significant effort reconstructing connectomes was controversial when we first started. One of its most outspoken proponents, Sebastian Seung, found himself at the center of a debate concerning the intellectual value and federal funding required to pay for the research and technology necessary to acquire connectomes. In contrast, there was considerable enthusiasm for functional modeling, but no consensus on what such a model would look like.
 |
Undaunted, I managed to identify a number of like-minded scientists and we concluded that, modulo our different ideas about what might constitute an interesting mesoscale functional model, the data necessary to infer such models would likely be available by 2019 for a model organism such as the fruit fly or larval-stage zebrafish and that we should lobby to ensure adequate funding to to achieve this goal. In the meantime, I had developed a scalable neural-network architecture for learning mesoscale models. The problem for me as an engineer at Google was what to do until 2019.
Our work in connectomics involves ten software engineers located in Zürich, Seattle and Mountain View. The resources we draw upon are expensive and, while Google has a history of taking on challenges whose expected short-term return on investment is uncertain, we try to make sure that we don't become involved in projects such that we have to rely upon primary sources that we can't control. Moreover, we think carefully about how to optimize our time to have the largest impact.
 |
And so I posed the following trade-off to the participants of the Kavli symposium: work on brains for ten years or work on AI for two years and then work on brains for five years and learn twice as much — and then I answered my question by stating that I believed my time would be best spent developing artificially intelligent systems to accelerate science itself.
Our connectomics work would not have been possible without what we've learned from the last decade of research on artificial neural networks. The recent focus on reinforcement learning has demonstrated that machines can learn tasks simply by interacting with the world, and the applications extend well beyond single-player video games and traditional two-player, complete-information games like Chess and Go.
 |
But these high-profile applications are just the tip of the iceberg. In addition to better understanding our existing tools in the process of solving banal problems, like captioning videos and improving speech recognition, we are developing prototype versions of digital assistants that use attention and related cognitive strategies to help users solve everyday problems.
 |
We've also seen a resurgence of interest in automated code synthesis and engineers have developed new tools that learn to optimize the design of neural network architectures. The most important accelerator for artificial intelligence in the next decade will be the recursive application of AI to building better AI systems. Just as Moore's Law accelerated the development of nearly everything having to do with computer technology, AI applied to hardware development will help to sustain Moore's Law while further accelerating AI technology.
 |
In this class, we propose to test the hypothesis that human-level AI is possible. From what well of confidence does such a conceit arise from? What do we really know about human intelligence? Here you see five books. The top two are the PDP books produced by Jay McClelland, David Rumelhart and the Parallel Distributed Processing (PDP) Research Group that set forth the basic concepts of connectionism, challenging the symbolic processing theory of cognition and redefining AI in terms of the massively parallel architecture of the human mind.
The other books are drawn from three separate sub disciplines of neuroscience: cognitive neuroscience, systems neuroscience and neurophysiology. These sub disciplines don't begin to cover the entire field, but each one provides a valuable perspective on biological computation and offers useful insight to engineers and computer scientists looking for ideas on how to build intelligent systems.
The PDP books leverage these insights, but then — unencumbered by biology, they explore new directions and offer new approaches for engineering intelligent systems. We haven't begun to exhaust any of these sources of inspiration, and have every reason to believe that cross fertilization between disciplines will continue to accelerate as technology for searching, digesting and integrating scientific literature improves.
 |
In order for you to understand where I'm coming from I'm going to tell you a little bit about my history. I started my graduate degree in computer science at Yale in 1982. I was super excited about everything having to do with computers and artificial intelligence. In addition to courses on compilers and operating systems, I took introductory courses in neuroscience to fulfill my PhD minor requirement.
I finished my PhD in 3 1/2 years with a thesis in artificial intelligence working with Drew McDermott and took on a position as an assistant professor at Brown University without really thinking about it. Most of the next 20 years are a blur as I wrote more than 100 papers and three books, was promoted to full professor, served as chair of the department for five years and then VP of IT and Deputy Provost of the University for three more.
In 2006, I needed a break frame from academia and took advantage of a year-long sabbatical to resurrect my interest in neuroscience. I found that what I had learned at Yale was woefully incomplete and much of it just wrong. In my last year as Provost I applied what I knew about probabilistic graphical models to implement a model of the cortex developed by David Mumford, a mathematician in the Applied Math department at Brown. I then accepted an invitation from Peter Norvig to spend my sabbatical at Google starting January 2006.
I had been a Google employee less than a week when I got an invitation from Dileep George and Jeff Hawkins to spend my sabbatical at Numenta, a startup Jeff was funding to conduct basic research in neuroscience. I was back at Google the following September and launched the Cortex project to explore the prospects for applying techniques from computational neuroscience to solving problems in machine vision.
After some initial missteps experimenting with restrictive Boltzmann machines we decided to ignore the advice about vanishing gradients and apply gradient descent to conventional multilayer perceptrons and convolutional neural networks. It became immediately clear that the bottleneck was training and so we focused on developing parallel algorithms and scaling numerical methods by increasing SIMD capacity in Google data centers.
We talked with Jeff Dean and Sanjay Ghemawat several times before Andrew Ng joined Google part-time and Jeff became the technical lead on what at the time was called DisBelief and would later become Google Brain. More than anything else it was Jeff's leadership and his ability to attract some of Google's best software engineers that led to the success of Google Brain, TensorFlow and the many technologies and products that these efforts made possible.
As mentioned earlier, the students taking CS379C in 2013 and I created a technology roadmap for scalable neuroscience, and, by February 2014, I had managed to tempt Viren Jain, then at the Howard Hughes Medical Institute Janelia Farm Research Campus to join Google and lead the Neuromancer Project to create the software that generated the connectomic reconstruction shown earlier. Alan Eustace who was then the SVP for Research encouraged us ramp up quickly, mentioning that Larry and Sergey were interested.
With Viren leading the Neuromancer team, I turned my attention to functional analysis, but quickly realized that it would be several years before scalable functional imaging would sufficiently advanced to handle whole-brain, pan-neuronal recording in adult behaving animals. My academic colleagues tried to convince me to work on smaller, more tractable circuits and were suspicious that my data-driven, machine-learning approach would not yield anything interesting by way of intelligible mesoscale models. Some believed my approach would fail because it didn't account for what we had learned about the brain, but I was suspicious about what we thought we knew, reluctant to use an impoverished possibly flawed prior and willing to let the data speak for itself. As a scientist, it is important to have a good sense of what we know and what we don't know2.
 |
Our goal in connectomics is to reconstruct all of the neurons in a tissue sample, but neurons are not the only type of cell found in the brain. There are approximately as many non-neuronal glial cells as there are neurons in the brain, classified unimaginatively as either macroglia or microglia. Macroglia include several specialized cell types including astrocytes that link neurons to the blood supply while maintaining the blood-brain barrier and oligodendrocytes that provide myelination to speed the transmission of electrical signals.
Microglia are considered the macrophages of the brain — they are responsible for clearing the brain of dead cells and related detritus. While still believed to perform an essential immunological function, we now think they play more complicated roles in both healthy and diseased brains. In the image shown in the upper left-hand corner of the slide, neurons are illuminated by green fluorescent protein indicators, the nuclei of neurons are indicated in blue and a number of microglial cells highlighted in red are visible closely tracking neuronal processes.
Microglia differentiate in the yolk sac and are perinatally exposed to many of the same factors that determine the function of the cells that comprise our innate immune system. They eventually migrate to the central nervous system where they become isolated by the blood brain barrier. They play a crucial role throughout development during which they prune developing neural circuits by reducing the number of neurons from approximately 200 billion to half that and the number of synapses from 200 trillion to 100 trillion. This winnowing takes place over several decades.
In the mature brain, they play a computational role related to learning that involves microglia using filopodia extensions to direct the formation of neural connections and influence synapse strength. Microglia make transient physical contact with all 100 trillion synapses of the mature brain every 30-60 minutes. The green arrows indicate the normal direction of microglia morphological changes with respect to the healthy brain and the orange arrows indicate that in some circumstances microglia can revert to earlier stages and damage cells mistaking by normal activity during learning as indications of dead or dying cells.
The video in the upper right shows microglia responding in their immunological role to local laser-induced damage by altering their shape to engulf and then digest damaged tissue. The video just below shows their normal behavior as they constantly survey nearby synapses. If the mother experiences an acute immune response while the immature microglia are still exposed to cells destined for the innate immune system, they can become epigenetically reprogrammed to misbehave in response to healthy neurons, resulting in the drastic pruning of dendrites we see in patients with schizophrenia, autism and related brain disorders.
Much of what I've told you has been discovered in the last decade. It almost seems we are in a negative-sum game in which the more we learn the more we realize we don't understand. I'm telling you this and the other examples in the earlier footnote to underscore the fact that the brain has not yielded its secrets easily. We desperately need better instruments to reveal the underlying physical processes and better analytical tools to make sense of the resulting rivers of data. The programmer's apprentice is a prototype for the neuroscientist's digital amanuensis.
P.S. You can learn about our efforts to learn more about microglia here, including a discussion of possible drug treatments to mitigate the neural damage inflicted when errant early programming causes microglia to attack healthy cells in adult brains.
1 The basic principles for electron microscopy and nuclear magnetic resonance imaging were discovered in the first half of the twentieth century, but it took decades to create the first useful instruments and decades more to improve the technology to operate anywhere near the scale of the modern technologies. It took the development of radar in the second world war to enable MRI technology. Materials science and the semiconductor industry were largely responsible for the advancement of electron microscopy to the stage where we can use it collect connectomic data for sizable tissue samples.
Maria Goeppert-Mayer explained the physics of two-photon absorption. The basic idea of two-photon-excitation microscopy was invented by Winfried Denk and James Strickler in 1990, but the development of useful two-photon microscopy for neural imaging would have to wait for better fluorescent dyes, new techniques for dealing with light scattering and exploiting properties of the point-spread function, and development of modern super-resolution imaging techniques.
2 The history of biology is riddled with misleading theories. Francis Crick's aptly named Central Dogma of molecular biology states that information in genes moves in one direction by way of a two-step process, transcription followed by translation: DNA to RNA to protein, but in subsequent decades multiple Nobel Prizes were awarded for work demonstrating this not to be an inviolable law, including Howard Temin and David Baltimore's independent discovery of Reverse Transcription.
Ernst Haeckel's produced drawings illustrating the different developmental stages of animals that he claimed supported his "ontogeny recapitulates phylogeny" Recapitulation Theory, still featured in many textbooks. This hypothesis is simply false. Camillo Golgi and Santiago Ramón y Cajal famously argued for decades before Cajal's neuroanatomical work decisively established the Neuron Doctrine, claiming that the nervous system is made up of discrete individual cells. But is it? What about gap junctions?
In 1999, Mario Galarreta working in Shaul Hestrin's lab at Stanford discovered a network of fast-spiking interneurons in the neocortex connected by electrical synapses, thereby establishing that electrical signal transmission via these so-called gap junctions plays an important role in adult organisms and was not merely a transition phase during early development as most scientists believed at the time. It wasn't easy convincing some of those who had an intellectual stake in the prevailing theory to adjust their world view.
The process of formulating hypotheses, running experiments and weighing the evidence is the foundation for the scientific method. I'm not suggesting it is fundamentally flawed, particularly slow or unusually biased. It's only as flawed, slow and biased as are human scientists. In his lecture in the 2015 instantiation of CS379C, Mario encouraged students to think about three categories of knowledge: the "Things we know", the "Things we know that we don't know" and the "Things that we don't know that we don't know" and consider if the distribution of these three types of knowledge in their chosen area of scientific inquiry is commensurate with their capacity for dealing with ambiguity and uncertainty.