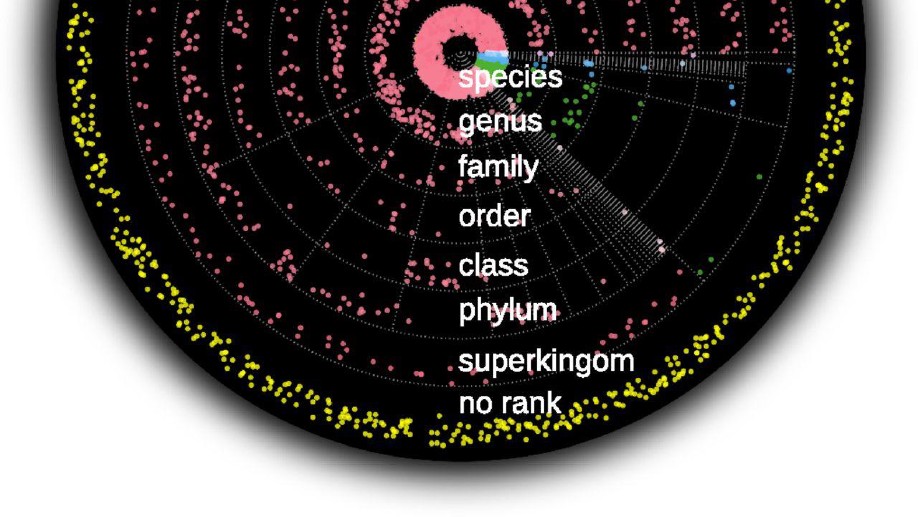
BACKGROUND: Plasma cell-free RNA (cfRNA) encompasses a broad spectrum of RNA species that can be derived from both human cells and …
Mark Kowarsky
PhD student
Stanford University
Biography
I’m a PhD student in the Quake lab applying machine learning and data visualization techniques to high-throughput sequencing data to answer questions across a range of biological fields. I have broad interests and a passion for thinking deeply about data to learn domain knowledge.
Biological experience includes: metagenomics, immunology, virology, marine biology, infectious diseases, developmental biology, microbiomes, and stem cell biology. Other interests include: quantum information and computational linguistics.
Interests
- Data Science
- Bioinformatics
- High-throughput sequencing
Education
-
PhD (expected) in Physics, 2019
Stanford University
-
MSc in Physics, 2012
University of Melbourne
-
BSc in Mathematical Physics, 2010
University of Melbourne