


Next:Estimating
:Up:Modelling
decisions for thePrevious:4.
Using prior populationContents
Allele frequency models
There are two basic models for the allele frequencies. One model assumes
that the allele frequencies in each population are independent draws from
a distribution that is specified by a parameter called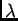 .
That is the original model that we used in
Pritchard
et al. 2000a . Usually we set
.
That is the original model that we used in
Pritchard
et al. 2000a . Usually we set 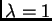 ;
this is the default setting. More recently, we have implemented a model
with correlated allele frequencies. This says that frequencies in the different
populations are likely to be similar (probably due to migration or shared
ancestry). Further details are given below.7
The independent model works well for many data sets. Roughly speaking,
this prior says that we expect allele frequencies in different populations
to be reasonably different from each other. The correlated frequencies
model says that they may actually be quite similar. This often improves
clustering for closely related populations, but may increase the risk of
over-estimating
;
this is the default setting. More recently, we have implemented a model
with correlated allele frequencies. This says that frequencies in the different
populations are likely to be similar (probably due to migration or shared
ancestry). Further details are given below.7
The independent model works well for many data sets. Roughly speaking,
this prior says that we expect allele frequencies in different populations
to be reasonably different from each other. The correlated frequencies
model says that they may actually be quite similar. This often improves
clustering for closely related populations, but may increase the risk of
over-estimating 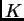 (see below). If one population is quite divergent from the others, the
correlated model can sometimes achieve better inference if that population
is removed.
(see below). If one population is quite divergent from the others, the
correlated model can sometimes achieve better inference if that population
is removed.
Subsections



Next:Estimating
:Up:Modelling
decisions for thePrevious:4.
Using prior populationContents
William Wen 2002-07-18