



Next: Multimodality
Up: Missing data, null alleles...
Previous: Dominant loci
Contents
The structure model assumes that loci are independent within
populations (i.e., not in LD within populations). This assumption is
likely to be violated for sequence data, or data from non-recombining
regions. The impact of using such data is likely to be that (1) the
algorithm underestimates the degree of uncertainty in ancestry
estimates, and in the worst case, may be biased or inaccurate; (2)
estimation of 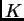 is unlikely to perform well.
One valid solution is to recode the haplotypes from a linked region so
that it is represented as a single locus with
is unlikely to perform well.
One valid solution is to recode the haplotypes from a linked region so
that it is represented as a single locus with  alleles. If there
are very many haplotypes, one could group related haplotypes together.
We are also aware of analyses that have taken the polymorphic sites
from sequence data from multiple regions, and treated these within
structure as independent loci. This type of analysis may yield
sensible and informative results, however considerable caution must be
applied to interpreting the results. The linkage model is likely to
perform better here than the independent sites model. We would not
recommend this type of approach for sequences from just one or a few
regions, except perhaps in purely exploratory analysis.
alleles. If there
are very many haplotypes, one could group related haplotypes together.
We are also aware of analyses that have taken the polymorphic sites
from sequence data from multiple regions, and treated these within
structure as independent loci. This type of analysis may yield
sensible and informative results, however considerable caution must be
applied to interpreting the results. The linkage model is likely to
perform better here than the independent sites model. We would not
recommend this type of approach for sequences from just one or a few
regions, except perhaps in purely exploratory analysis.




Next: Multimodality
Up: Missing data, null alleles...
Previous: Dominant loci
Contents
William Wen
2002-07-18