
Our laboratory is interested in understanding core molecular metabolism and physiology of prokaryotic microorganisms involved in environmental, bioenergy, and intestinal processes.
In particular, we focus on two areas:
RESEARCH AREA 1: Molecular mechanisms of electron flow in energy conservation of key anaerobic microorganisms
Life is redox chemistry and inherently coupled to energy-conserving oxidation/reduction reactions. In contrast to well-characterized proteobacteria, electron flow and associated energy conservation follows different, evolutionarily more ancient principles in strictly anaerobic microorganisms, such as methanogens, homoacetogens, and reductively dehalogenating microorganisms.
Project 1: Electron flow through the electron-bifurcating Vhu/Fdh-Hdr supercomplex
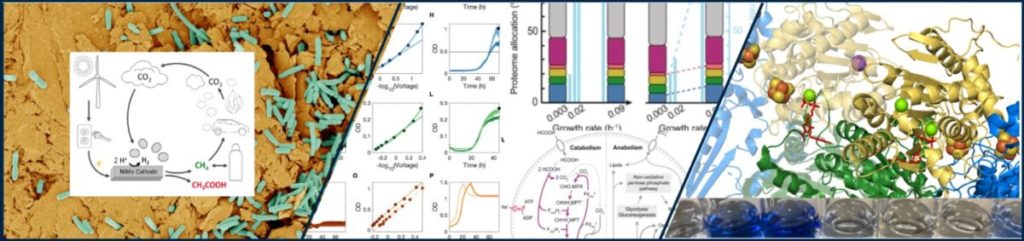
In hydrogenotrophic methanogens, the Vhu/Fdh-Hdr complex catalyzes the bifurcation of H2-derived electrons as a key step in energy conservation. This novel, flavin-dependent electron bifurcation couples the endergonic reducing of CO2 to formylmethanofuran with the exergonic reduction of the heterodisulfide (CoM-S-S-CoB). We are investigating electron flow from hydrogenases and formate dehydrogenases through Vhu/Fdh-Hdr to CoM-S-S-CoB and formylmethanofuran dehydrogenase (ferredoxin). We also discovered that the hydrogenases and the Vhu/Fdh-Hdr supercomplex show extraordinary stability when coupled to an electrochemical platform to initiate electron flow through the enzyme. We are conducting structure-function analyzes to deconvolute the unusual catalytic and stability properties of these enzymes, and explore new platforms for how these enzymes and microorganisms can be used in bio-electrochemical systems in bioenergy.

Project 2: Uptake and metabolism of cathode-derived electrons
Direct electron transfer between microorganisms as well as between a microbe and a redox-active surface is frequent and plays important roles in geochemical as well as intestinal and pathogenic environments. The molecular determinants required for direct uptake of electrons seems to be different from those involved in electron release, e.g. to humic acids or other insoluble electron carriers. We found that the same molecular elements involved in direct cathodic electron uptake are also involved in corrosion of Fe0. We are studying the sulfate-reducing ‘Desulphopila corrodens’ IS4 and the genetically tractable Desulfovibrio ferrophilus IS5 in molecular detail.

Project 3: Mechanism of energy conservation by reductive dehalogenation
Chloroethenes, such as tetrachloroethene (PCE) and trichlorethene (TCE) are the most prevalent groundwater contaminants in the developed world, and clean-up costs are estimated to be billions of dollars. Bioremediation of these chlorinated ethenes to harmless ethene is a stepwise, strictly anaerobic process where each chlorine atom is sequentially eliminated reductively as Cl–. Most important is microbial catabolic reductive dehalogenation using H2 as electron donor. Complete reductive dehalogenation of PCE or TCE is mediated by two physiologically distinguishable groups of microorganisms. Group one consists of phylogenetically diverse anaerobes, such as Desulfitobacterium species but also Dehalococcoides sp, and typically mediate dehalogenation to cDCE. The second group consists so far only of Dehalococcoides (Dhc) species, which are phylogenetically deeply branching Chloroflexi, that mediate reduction of c-DCE and vinyl chloride to ethene.
Specifically, we are investigating the enzymology and maturation of reductive dehalogenases of the Dhc type, the diversity and ecophysiological function of reductive dehalogenases in non-contaminated environments, and the evolution of reductive dehalogenases and dehalogenating microorganisms.

Project 4: Metabolic processes and microbial interactions in human intestinal microbial communities
Irritable Bowel Syndrome (IBS) is a chronic, episodic gastrointestinal disorder that is characterized by abdominal pain and altered bowel habits. IBS prevalence is estimated to be 10-15% in Western countries comprising 25 to 50 percent of all referrals to gastroenterologists. The gastrointestinal tract harbors a complex and diverse microbial community, which plays important roles in host nutrition, immune function, health and disease, and it is hypothesized the IBS disease phenotype is associated with a change in colonic microbiota and/or host factors such as mucosal function and immunity. This is strengthened by reports that luminal antibiotics or probiotic treatment may be effective in alleviating symptoms in patients with IBS. Indirect evidence for alterations in the microflora of humans with IBS comes from changes in colonic fermentation patterns has been described in patients with IBS.
Specifically, we are investigating the structure of microbial populations and communities associated with IBS, as well as the metabolic interactions between gut microorganisms, and how these interactions can be used to develop cures for the disease.
RESEARCH AREA 2: Control of energy and resource allocation to cellular processes under slow growth conditions
In this research area, we seek to understand how microorganisms change allocation of cellular resources under conditions of slow or no growth. In other words, how do cells coordinate catabolism and anabolism in the spatially constrained cellular proteome, when the demand for cellular energy and biosynthetic capacity changes? With decreasing growth rate, enteric bacteria appear to switch metabolism from an inefficient fermentative metabolism to a more efficient O2 respiratory metabolism. This seem to reflect that a smaller fermentative proteome enables a larger flux of ATP than the more efficient respiratory proteome. However, methanogens and homoacetogenic bacteria such as the deepsea sediment Chloroflexi inherently cannot adapt to slow growth by splitting metabolism. We study the underlying features and mechanisms in chemostat cultures using proteomic, transcriptomic and metabolomic approaches in conjunction with genetics.
Project 5: Genomic reconstruction of energy metabolism of deepsea sediment microorganisms
Most microorganisms in deepsea sediments are in a low flux, energy-limited environment, where their estimated doubling times are in the range of decades rather than minutes or hours. How do microbes acclimate to such conditions, and what are the molecular and physiological features of such slow growth state, where maintenance rather than growing is the predominant mode of living? In order to uncover the underlying mechanism(s), we focus on marine Chloroflexi, which are highly enriched in many deepsea sediments. Using single cell genomics and binned metagenomes, we are reconstructing complete metabolic maps for energy conservation in these Chloroflexi. One preeminent feature we have been discovering is the predominance to the Wood Ljungdahl pathway in many but not all deepsea Chloroflexi. Current efforts focus on relating pathway structure and combinations to fitness in such low flux environments. Moreover, these molecular insights have been providing the basis for novel enrichment strategies of Chloroflexi from deepsea sediments to test our hypothesis.
