 |
Arenaviridae Biology |
Basics
Arenaviruses are single stranded, negative sense RNA viruses with two nucleic acid segments, one segment being slightly larger in the overall 5000-7400 base pair genome. Virions are enveloped, pleomorphic sperical structures with distinct, club-shaped surface projections. Host ribosomes within the viral envelope give the virus its characteristic sandy appearance. Nucleocapsid structure is helical.
Arenavirus Taxonomy
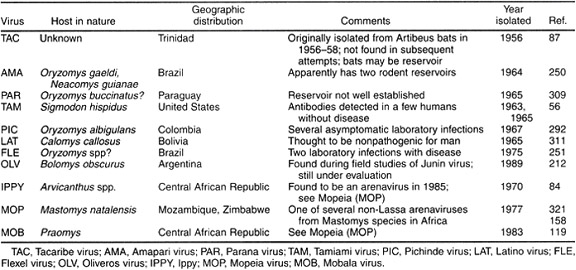
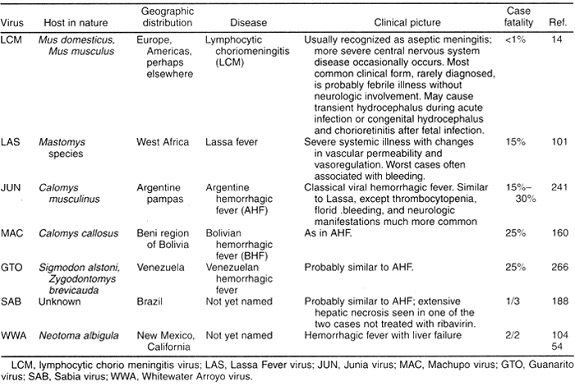
The virus family Arenaviridae consists of only one genus, yet within this genus, most viruses are divided into two different clades: the Old World arenaviruses, and the New World arenaviruses (also known as the Tacaribe complex). The differences between the two groups have been established through the use of serologic assays. Epitopes shared by Old World and New World arenaviruses have been detected on NP and GP2 using monoclonal antibodies. Additionally, the New World viruses have been grouped even more specifically into three lineages based on antigenic data: two that are pathogenic in humans, A and B, and one that is not, C. The following lists of lineages reference only those arenaviruses that are pathogenic in humans, though it should be noted that each lineage contains some viruses that are non-pathogenic. Tacaribe complex lineage B consists of Junin, Guaranito, Sabia, and Machupo viruses, which appear to produce similar symptoms in humans, though they are genetically distinct from one another. The universally highly pathogenic effect of infection with this lineage of viruses suggests a radiational evolution from a common ancestor. Tacaribe complex lineage A consists of White Water Arroyo and Pirital viruses, which are also antigenically similar. With regards to the Old World arenaviruses, antigenic factors differentiate Lassa virus from Lymphocytic choriomeningitis virus, which is the Old World arenavirus that is most similar to New World arenaviruses. However, these clades and lineages are malleable and subject to change as new genetic information becomes available. For instance, analysis of glycoprotein sequences suggest that two New World arenaviruses, Pichinde and Oliveros, are in fact more similar to Old World arenaviruses than their New World counterparts. An interesting study might compare Pichinde and Oliveros RNA to that of lymphocytic choriomeningitis virus. Through increasingly specific analysis of arenavirus genetic material and proteins, researchers are gaining a clearer picture of the evolution of arenaviruses.
Tables courtesy of Fields Online:
Arenaviruses Pathogenic in Humans
Arenaviruses Non-Pathogenic in Humans
http://gateway.ut.ovid.com.laneproxy.stanford.edu/gw1/ovidweb.cgi
Genetics of Arenaviruses
As a byproduct of their segmentation, genetic reassortment of the S(mall) and L(arge) genomic RNAs within progeny viruses following simultaneous infection with parental virus strains, has been observed in the laboratory between wild-type and temperature-sensitive mutants of Pichinde virus. However, naturally occurring LCMV reassortments have been recovered after dual infections with two wild-type parental strains. LCMV also exhibits diploidy, the property of having multiple genomic “alleles” packaged in the same virion. In one case, a new strain of LCMV contained the L RNA of one parent strain, and the S RNA’s of both parents. Additionally, A Lassa-Mopeia reassortant virus, containing the L RNA of Mopeia and the S RNA of Lassa, has been developed in the lab, and was in fact found to have significantly diminished mortality in suckling mice. This may suggest vaccine potential.