Cross-Correlation
We developed an unbiased, automated and easily implemented assay to characterize asymmetric subcellular protein and organelle localization within a tissue. Specifically, the cross-correlation score was monitored upon lateral displacement of one channel. As shown below, an artificial overlap of membrane and asymmetric fluorescence at the posterior side of the cells was caused by the shift.
To analyze whether these relative peaks in cross-correlation were significant, we performed two separate controls. First, we randomized the position of individual pixels within each row of the asymmetric channel (actin) (Fig. 1B). As a second control, we took the cross-correlation analysis data obtained in Fig. 2 (top left) and randomized the position of individual pixels within each row of the heatmap (Fig. 1C).

We suggest setting a threshold of 4 standard deviations (4 sigma) above the mean value to provide a high likelihood of significance. Assuming a normal distribution, the probability to reach by chance an average cross-correlation score that is above this threshold is less than 0.01% (p<10-4).
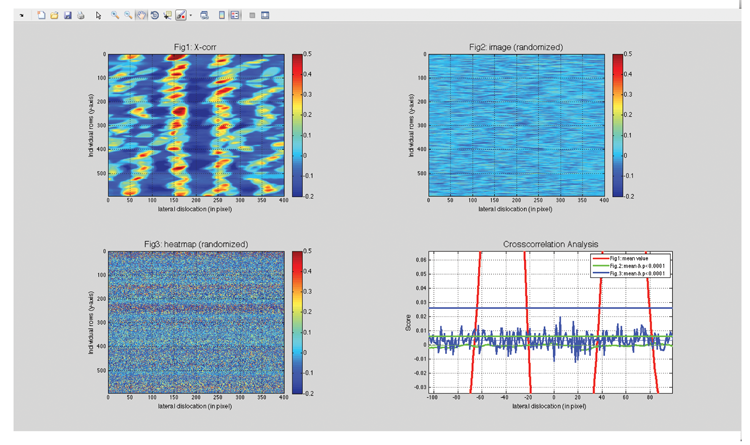
We developed a Matlab script of this analysis (screenshot of output shown below). The top left panel shows the cross-correlation of the two chanels analyzed. The two randomization controls are shown in the top right panel (randomized image) and bottom left panel (randomized heatmap), respectively. Average crosscorrelation values for all three images are summarized in the bottom right panel.
To download the annotated, fully automated Matlab script performing the analysis, as well as sample images, please click here. The link to the PubMed citation for this work is found here.