Models for binary outcomes¶
Data \((X_i, Y_i), 1 \leq i \leq n\).
Examples:
medical: presence or absence of cancer
financial: bankrupt or solvent
industrial: passes a quality control test or not
For \(0-1\) responses we need to model $\(\pi(x_1, \dots, x_p) = P(Y=1|X_1=x_1,\dots, X_p=x_p)\)$
options(repr.plot.width=5, repr.plot.height=4)
Binary outcomes¶
Likelihood (\(X\) considered fixed or likelihood considered conditional) $\( -\log L(\pi|Y) = \sum_{i=1}^n - Y_i \log\left(\frac{\pi_i}{1-\pi_i}\right) - \log(1 -\pi_i) \)$
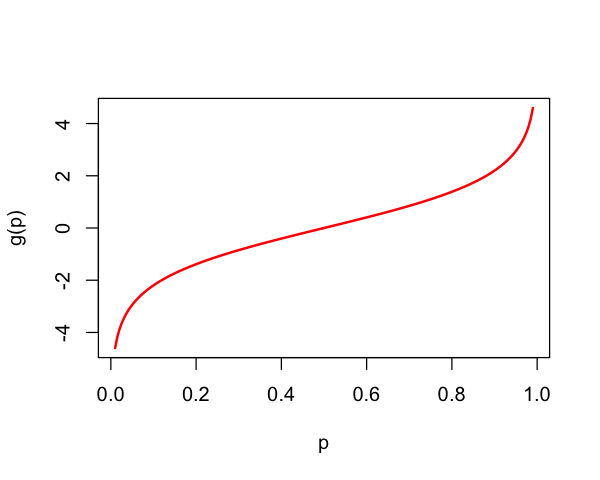
The function $\( \pi \mapsto \log\left(\frac{\pi}{1-\pi}\right) \overset{\text{def}}{=} \text{logit}(\pi) \)$
Logit transform¶
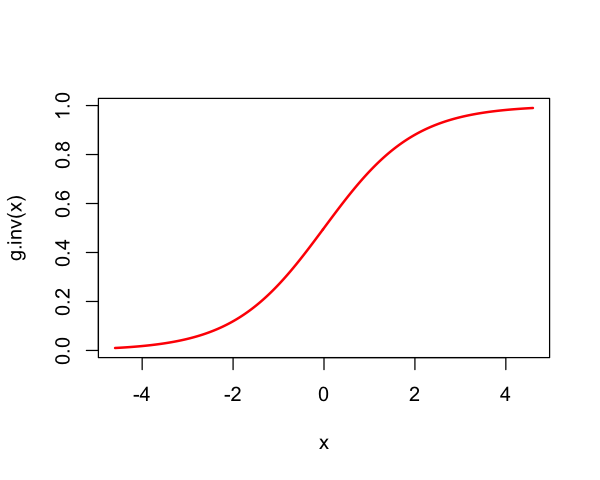
Note $\( \text{logit}\left(\frac{e^{\eta}}{1+e^{\eta}}\right) = \eta. \)\( where \)\( F_{\text{logistic}}(\eta) = \frac{e^{\eta}}{1+e^{\eta}} \)$ is the logistic distribution function.
So, $\( \text{logit} = F_{\text{logistic}}^{-1} \)$
g = function(p) {
return(log(p / (1 - p)))
}
p = seq(0.01,0.99,length=200)
plot(p, g(p), lwd=2, type='l', col='red')
g.inv <- function(x) {
return(exp(x) / (1 + exp(x)))
}
x = seq(g(0.01), g(0.99), length=200)
plot(x, g.inv(x), lwd=2, type='l', col='red')
Logistic regression model¶
Write likelihood in natural parameters
Then,
Logistic regression model: \(\eta_i=X_i^T\beta\) and
Logistic regression¶
In other words,$\( \text{logit}(\pi(x_1,\dots,x_p)) = x^T\beta. \)$
Negative log-likelihood is convex in \(\beta\) \(\implies\) easy to add regularization penalties (and solve such problems…)
Default in most software for GLMs because(?) \(\eta_i=X_i^T\beta\) (i.e. canonical link)
Logistic regression¶
How do we know it’s convex?
Negative log-likelihood of saturated model with parameters \(\eta=(\eta_1, \dots, \eta_n)\)
with \(\Lambda_B(\eta)\) the cumulant generating function (CGF) of Bernoulli
For exponential family with sufficient statistic \(W\) and natural parameters \(\eta\)
where \(\mathbb{P}_0\) is the law under \(\eta=0\). In our case
Negative log-likelihood is $\( \eta \mapsto -\eta^TY + \Lambda(\eta) \)$
Logistic regression imposes linear constraint on \(\eta\): \(\eta \in \text{col}(X)\).
Logistic regression¶
Logistic is “natural” because it is a linear model for natural parameters of Binomial data.
It can appear in other places (c.f. 5.1.5 of Agresti)
Suppose we observe observations from $\( \begin{aligned} Y &\sim \text{Bernoulli}(\pi) \\ X | Y \sim & \begin{cases} N(\mu_0, \Sigma) & Y = 0\\ N(\mu_1, \Sigma) & Y=1 \end{cases} \end{aligned} \)$
Then, logistic model for \(Y|X\) holds.
Another reason this model is popular relates to the odds ratio (c.f. 5.1.4 of Agresti)
If the logistic model is correct along with an assumption about sampling, it is possible to estimate parameters of \(Y|X\) distribution in case-control studies where the actual randomness is \(X|Y\).
This is similar to (2.2.6) of Agresti (Lung Cancer and Smoking) which we discussed earlier.
Interpretation of coefficients: Odds Ratios¶
Logistic model:
If \(X_j \in \{0, 1\}\) is dichotomous, then odds for group with \(X_j = 1\) are \(e^{\beta_j}\) higher, other parameters being equal.
Rare disease hypothesis¶
When incidence is rare, \(P(Y=0)\approxeq 1\) no matter what the covariates \(X_j\)’s are.
In this case, odds ratios are almost ratios of probabilities: $\(OR_{X_j} \approxeq \frac{{\mathbb{P}}(Y=1|\dots, X_j=x_j+1, \dots)}{{\mathbb{P}}(Y=1|\dots, X_j=x_j, \dots)}\)$
Hypothetical example: in a lung cancer study, if \(X_j\) is an indicator of smoking or not, a \(\beta_j\) of 5 means for smoking vs. non-smoking means smokers are \(e^5 \approx 150\) times more likely to develop lung cancer
Other (linear) binary models¶
Suppose there is some latent variable \(T_i | X_i \sim F\) such that
Then,
Example: probit \(F=\Phi\) (dbn of \(N(0,1)\))
Setting $\( \eta_i = \text{logit}(F(X_i^T\beta)) \)\( the likelihood is then \)\( -\log L(\beta|Y) = \sum_{i=1}^n -Y_i \cdot \text{logit}(F(X_i^T\beta)) - \log(1 - F(X_i^T\beta)). \)$
Parameters no longer interpretable in odds ratios but problem can still be convex (e.g. probit)
Modelling probabilities¶
Note: \(\text{Var}(Y) = \pi ( 1 - \pi) = E(Y) \cdot ( 1- E(Y))\)
The binary nature forces a relation between mean and variance of \(Y\).
This makes logistic regression a Generalized Linear Model, use
glminR.
Horseshoe crabs¶
library(glmbb) # for `crabs` data
data(crabs)
head(crabs)
| color | spine | width | satell | weight | y | |
|---|---|---|---|---|---|---|
| <fct> | <fct> | <dbl> | <int> | <int> | <int> | |
| 1 | medium | bad | 28.3 | 8 | 3050 | 1 |
| 2 | dark | bad | 22.5 | 0 | 1550 | 0 |
| 3 | light | good | 26.0 | 9 | 2300 | 1 |
| 4 | dark | bad | 24.8 | 0 | 2100 | 0 |
| 5 | dark | bad | 26.0 | 4 | 2600 | 1 |
| 6 | medium | bad | 23.8 | 0 | 2100 | 0 |
width_glm = glm(y ~ width, family=binomial(), data=crabs)
summary(width_glm)
anova(glm(y ~ 1, family=binomial(), data=crabs), width_glm)
c(225.76-194.45, 4.887^2)
Call:
glm(formula = y ~ width, family = binomial(), data = crabs)
Deviance Residuals:
Min 1Q Median 3Q Max
-2.0281 -1.0458 0.5480 0.9066 1.6942
Coefficients:
Estimate Std. Error z value Pr(>|z|)
(Intercept) -12.3508 2.6287 -4.698 2.62e-06 ***
width 0.4972 0.1017 4.887 1.02e-06 ***
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
(Dispersion parameter for binomial family taken to be 1)
Null deviance: 225.76 on 172 degrees of freedom
Residual deviance: 194.45 on 171 degrees of freedom
AIC: 198.45
Number of Fisher Scoring iterations: 4
| Resid. Df | Resid. Dev | Df | Deviance | |
|---|---|---|---|---|
| <dbl> | <dbl> | <dbl> | <dbl> | |
| 1 | 172 | 225.7585 | NA | NA |
| 2 | 171 | 194.4527 | 1 | 31.30586 |
- 31.31
- 23.882769
summary(glm(y ~ width, data=crabs)) # same fit as `lm`
Call:
glm(formula = y ~ width, data = crabs)
Deviance Residuals:
Min 1Q Median 3Q Max
-0.8614 -0.4495 0.1569 0.3674 0.7061
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -1.76553 0.42136 -4.190 4.46e-05 ***
width 0.09153 0.01597 5.731 4.42e-08 ***
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
(Dispersion parameter for gaussian family taken to be 0.1951497)
Null deviance: 39.780 on 172 degrees of freedom
Residual deviance: 33.371 on 171 degrees of freedom
AIC: 212.26
Number of Fisher Scoring iterations: 2
summary(lm(y ~ width, data=crabs)) # dispersion is \hat{\sigma}^2
0.4418^2
Call:
lm(formula = y ~ width, data = crabs)
Residuals:
Min 1Q Median 3Q Max
-0.8614 -0.4495 0.1569 0.3674 0.7061
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -1.76553 0.42136 -4.190 4.46e-05 ***
width 0.09153 0.01597 5.731 4.42e-08 ***
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Residual standard error: 0.4418 on 171 degrees of freedom
Multiple R-squared: 0.1611, Adjusted R-squared: 0.1562
F-statistic: 32.85 on 1 and 171 DF, p-value: 4.415e-08
Logistic regression and \(I \times 2\) tables¶
For \(I \times 2\) tables \(N=(N_{ij})\), set
Under independent rows sampling,
Under Poisson sampling, same result holds if we condition on \(N_{i+}\).
Model
Like a one-way ANOVA regression model
Need some identifiability constraint (
Roften sets one \(\beta\) to 0).Not really using a logistic assumption here – just parameterizing \(\pi_i\) a certain way: model is saturated.
# Table 3.8
absent = c(17066, 14464, 788, 126, 37)
present = c(48, 38, 5, 1, 1)
total = absent + present
scores = c(0, 0.5, 1.5, 4.0, 7.0)
proportion = present / total
alcohol = rbind(absent, present)
chisq.test(alcohol)
Warning message in chisq.test(alcohol):
“Chi-squared approximation may be incorrect”
Pearson's Chi-squared test
data: alcohol
X-squared = 12.082, df = 4, p-value = 0.01675
lr_stat = function(data_table) {
chisq_test = chisq.test(data_table)
return(2 * sum(data_table * log(data_table / chisq_test$expected)))
}
lr_stat(alcohol)
Warning message in chisq.test(data_table):
“Chi-squared approximation may be incorrect”
alcohol_scores = glm(proportion ~ factor(scores), family=binomial(),
weights=total)
summary(alcohol_scores)
Call:
glm(formula = proportion ~ factor(scores), family = binomial(),
weights = total)
Deviance Residuals:
[1] 0 0 0 0 0
Coefficients:
Estimate Std. Error z value Pr(>|z|)
(Intercept) -5.87364 0.14454 -40.637 <2e-16 ***
factor(scores)0.5 -0.06819 0.21743 -0.314 0.7538
factor(scores)1.5 0.81358 0.47134 1.726 0.0843 .
factor(scores)4 1.03736 1.01431 1.023 0.3064
factor(scores)7 2.26272 1.02368 2.210 0.0271 *
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
(Dispersion parameter for binomial family taken to be 1)
Null deviance: 6.2020e+00 on 4 degrees of freedom
Residual deviance: 1.8163e-13 on 0 degrees of freedom
AIC: 28.627
Number of Fisher Scoring iterations: 4
anova(glm(proportion ~ 1, family=binomial(), weights=total),
glm(proportion ~ factor(scores), family=binomial(), weights=total))
| Resid. Df | Resid. Dev | Df | Deviance | |
|---|---|---|---|---|
| <dbl> | <dbl> | <dbl> | <dbl> | |
| 1 | 4 | 6.201998e+00 | NA | NA |
| 2 | 0 | 1.816325e-13 | 4 | 6.201998 |
A 1 df test¶
In this case,
scoresis ordered, so we might fit model $\( \text{logit}(\pi_i) = \alpha + \beta \cdot \text{scores}_i. \)$Perhaps more power because direction of deviation from null is specified.
summary(glm(proportion ~ scores, family=binomial(), weights=total))
Call:
glm(formula = proportion ~ scores, family = binomial(), weights = total)
Deviance Residuals:
1 2 3 4 5
0.5921 -0.8801 0.8865 -0.1449 0.1291
Coefficients:
Estimate Std. Error z value Pr(>|z|)
(Intercept) -5.9605 0.1154 -51.637 <2e-16 ***
scores 0.3166 0.1254 2.523 0.0116 *
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
(Dispersion parameter for binomial family taken to be 1)
Null deviance: 6.2020 on 4 degrees of freedom
Residual deviance: 1.9487 on 3 degrees of freedom
AIC: 24.576
Number of Fisher Scoring iterations: 4
anova(glm(proportion ~ 1, family=binomial(), weights=total),
glm(proportion ~ scores, family=binomial(), weights=total)) # LR test
data.frame(df.1=pchisq(4.25, 1, lower.tail=FALSE),
df.4=pchisq(6.2, 4, lower.tail=FALSE))
| Resid. Df | Resid. Dev | Df | Deviance | |
|---|---|---|---|---|
| <dbl> | <dbl> | <dbl> | <dbl> | |
| 1 | 4 | 6.201998 | NA | NA |
| 2 | 3 | 1.948721 | 1 | 4.253277 |
| df.1 | df.4 |
|---|---|
| <dbl> | <dbl> |
| 0.03925033 | 0.1847017 |
Deviance¶
Instead of sum of squares, logistic regression uses deviance:
Defined as
where \(\pi\) is a location estimator for \(Y\) with \(\hat{\pi}_S\) the saturated fit (i.e. no contraints on \(\pi\)’s).
For Binomial data \(Y_i \sim \text{Binom}({\tt total}_i,\pi_i)\)
If \(Y\) is a binary vector, with mean vector \(\pi\) then
If \(Y_i \sim \text{Binom}({\tt total}_i,\pi_i)\) then
with \(\hat{\pi}_{S,i} = Y_i / {\tt total}_i\) is the MLE under the saturated model.
Fit using
weights=totalinglm.
Deviance for logistic regression¶
For any binary regression model, the deviance is:
For the logistic model, the RHS is:
The logistic model is special in that \(\text{logit}(\pi_i(\beta))=X_i^T\beta\).
Fitting the logistic model¶
Derivatives of log-likelihood¶
Let \(Y \in \{0,1\}^n, X \in \mathbb{R}^{n \times p}\).
Gradient / Score
Hessian
Above,
Derivatives related to moments as this is an exponential family (conditional on \(X\)) with natural parameter \(\beta\), sufficient statistic \(X^TY\) and cumulant generating function
Newton-Raphson¶
Given initial \(\hat{\beta}^{(0)}\)
Until convergence $\( \hat{\beta}^{(i+1)} = \hat{\beta}^{(i)} - (X^T W_{\hat{\beta}^{(i)}}(X)X)^{-1}X^T(Y - \pi_{\hat{\beta}^{(i)}}(X)) \)$
Inference¶
Suppose model is true, with true parameter \(\beta^*\). Then, $\( E_{\beta^*}[X^T(Y - \pi_{\beta^*}(X))] = 0 \)$
Taylor series $\( \pi_{\hat{\beta}}(X) \approx \pi_{\beta^*}(X) + W_{\beta^*}(X) X(\hat{\beta}-\beta^*) \)$
How big is remainder?
Inference¶
Stationarity conditions then read $\( X^T(Y - E_{\beta^*}[Y|X]) \approx X^TW_{\beta^*}(X) X(\hat{\beta}-\beta^*) \)$
Or, $\( \begin{aligned} \hat{\beta}-\beta^* &\approx \left(X^TW_{\beta^*}(X) X\right)^{-1}X^T(Y - E_{\beta^*}[Y|X]) \\ &= (X^TW_{\beta^*}(X)X)^{-1}X^T(Y-\pi_{\beta^*}(X)) \end{aligned} \)$
Implies $\( \begin{aligned} \text{Var}_{\beta^*}(\hat{\beta}-\beta^*) &\approx \left(X^TW_{\beta}^*(X) X\right)^{-1} \text{Var}_{\beta^*}\left(X^T(Y - \pi_{\beta^*}(X))\right) \left(X^TW_{\beta^*}(X) X\right)^{-1} \\ &\approx \left(X^T W_{\beta^*}(X) X\right)^{-1} \\ \end{aligned} \)$ (Last line assumes the model is true.)
Leads to
(Plugin estimator of variance appeals to Slutsky, consistency of \(\hat{\beta}\).)
Where do we really use the model?¶
Under the logistic model$\( \begin{aligned} \text{Var}_{\beta^*}(X^T(Y-\pi_{\beta}^*(X)) & \approx X^TW_{\beta^*}(X)X. \end{aligned} \)$
Really just using the standard asymptotic distribution of the MLE:
For logistic regression, note that (conditional on \(X\)) $\( E_{\beta^*}[\nabla^2 (-\log L(\beta^*|Y))] = \nabla^2 (-\log L(\beta^*|Y)). \)$
center = coef(width_glm)['width']
SE = sqrt(vcov(width_glm)['width', 'width'])
U = center + SE * qnorm(0.975)
L = center + SE * qnorm(0.025)
data.frame(L, center, U)
| L | center | U | |
|---|---|---|---|
| <dbl> | <dbl> | <dbl> | |
| width | 0.2978326 | 0.4972306 | 0.6966286 |
These are slightly different from what R will give you if you ask it for
confidence intervals:
confint(width_glm)
Waiting for profiling to be done...
| 2.5 % | 97.5 % | |
|---|---|---|
| (Intercept) | -17.8100090 | -7.4572470 |
| width | 0.3083806 | 0.7090167 |
What is logistic regression even estimating?¶
Before appealing to general properties of MLE, we had only really used the assumption the model was true once.
What can we say if the model is not true?
Suppose \((Y_i,X_i) \overset{IID}{\sim} F, 1 \leq i \leq n\).
Define the population parameter \(\beta^*=\beta^*(F)\) by $\( E_F[X(Y - \pi_{\beta^*(F)}(X))] = 0. \)$
This is minimizer of
Taylor expansion yields
Note that (under conditions…)
Implies $\( \hat{\beta}-\beta^*(F) \overset{D}{\to} N(0, n^{-1} Q(F)^{-1}\Sigma(F)Q(F)^{-1}). \)$
Variance above has so-called sandwich form.
Applying Slutsky $\( \hat{\beta}-\beta^*(F) \approx N\left(0, (X^TW_{\hat{\beta}}(X) X)^{-1} (n\Sigma(F)) (X^TW_{\hat{\beta}}(X) X)^{-1}\right) \)$
We know everything in the approximation above (or can estimate it) except \(\Sigma(F)\)
Covariance \(\Sigma(F)\) can be estimated, say, by bootstrap or sandwich estimator. (Approachable description by D. Freedman here).
Alternatively, entire logistic regression could be bootstrapped.
Testing in logistic regression¶
What about comparing full and reduced model?
For a model \({\cal M}\), \(DEV({\cal M})\) replaces \(SSE({\cal M})\).
In least squares regression, we use
This is replaced with (when model true)
These tests assume model \({\cal M}_F\) is correctly specified. Otherwise, Wald-type test with sandwich or other covariance estimate would have to be used.
Deviance tests do not agree numerically with Wald tests even when model is correct. Both are often used. Wald can fail!
bigger_glm = glm(y ~ width + weight, data=crabs, family=binomial())
anova(width_glm, bigger_glm)
| Resid. Df | Resid. Dev | Df | Deviance | |
|---|---|---|---|---|
| <dbl> | <dbl> | <dbl> | <dbl> | |
| 1 | 171 | 194.4527 | NA | NA |
| 2 | 170 | 192.8919 | 1 | 1.560777 |
We should compare this difference in deviance with a \(\chi^2_1\) random variable.
1 - pchisq(1.56,1)
Let’s compare this with the Wald test:
summary(bigger_glm)
Call:
glm(formula = y ~ width + weight, family = binomial(), data = crabs)
Deviance Residuals:
Min 1Q Median 3Q Max
-2.1127 -1.0344 0.5304 0.9006 1.7207
Coefficients:
Estimate Std. Error z value Pr(>|z|)
(Intercept) -9.3547261 3.5280465 -2.652 0.00801 **
width 0.3067892 0.1819473 1.686 0.09177 .
weight 0.0008338 0.0006716 1.241 0.21445
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
(Dispersion parameter for binomial family taken to be 1)
Null deviance: 225.76 on 172 degrees of freedom
Residual deviance: 192.89 on 170 degrees of freedom
AIC: 198.89
Number of Fisher Scoring iterations: 4