Clustering#
Download#
Clustering#

As in classification, we assign a class to each sample in the data matrix. However, the class is not an output variable; we only use input variables.
Clustering is an unsupervised procedure, whose goal is to find homogeneous subgroups among the observations.
Basic approaches#
Divisive / partitional: prototype being \(K\)-means clustering
Agglomerative: the prototype being hierarchical clustering
\(K\)-means clustering#
\(K\) is the number of clusters and must be fixed in advance.
The goal of this method is to maximize the similarity of samples within each cluster:

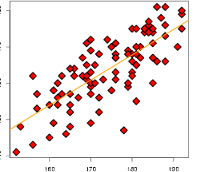
Without scaling#
With scaling#
\(K\)-means clustering algorithm#
Assign each sample to a cluster from 1 to \(K\) arbitrarily, e.g. at random (or smarter
kmeans++)Iterate these two steps until the clustering is constant:
Find the centroid of each cluster \(C_\ell\); i.e. the average \(\overline x_{\ell,:}\) of all the samples in the cluster:
\[x_{\ell,j} = \frac{1}{|C_\ell|}\sum_{i\in C_\ell} x_{i,j} \quad \text{for }j=1,\dots,p.\]Reassign each sample to the nearest centroid.

The algorithm is a descent algorithm – it terminates when no gain can be found by moving any point to any other cluster.
Each initialization could yield a different minimum.
\(K\)-means output with different initializations#
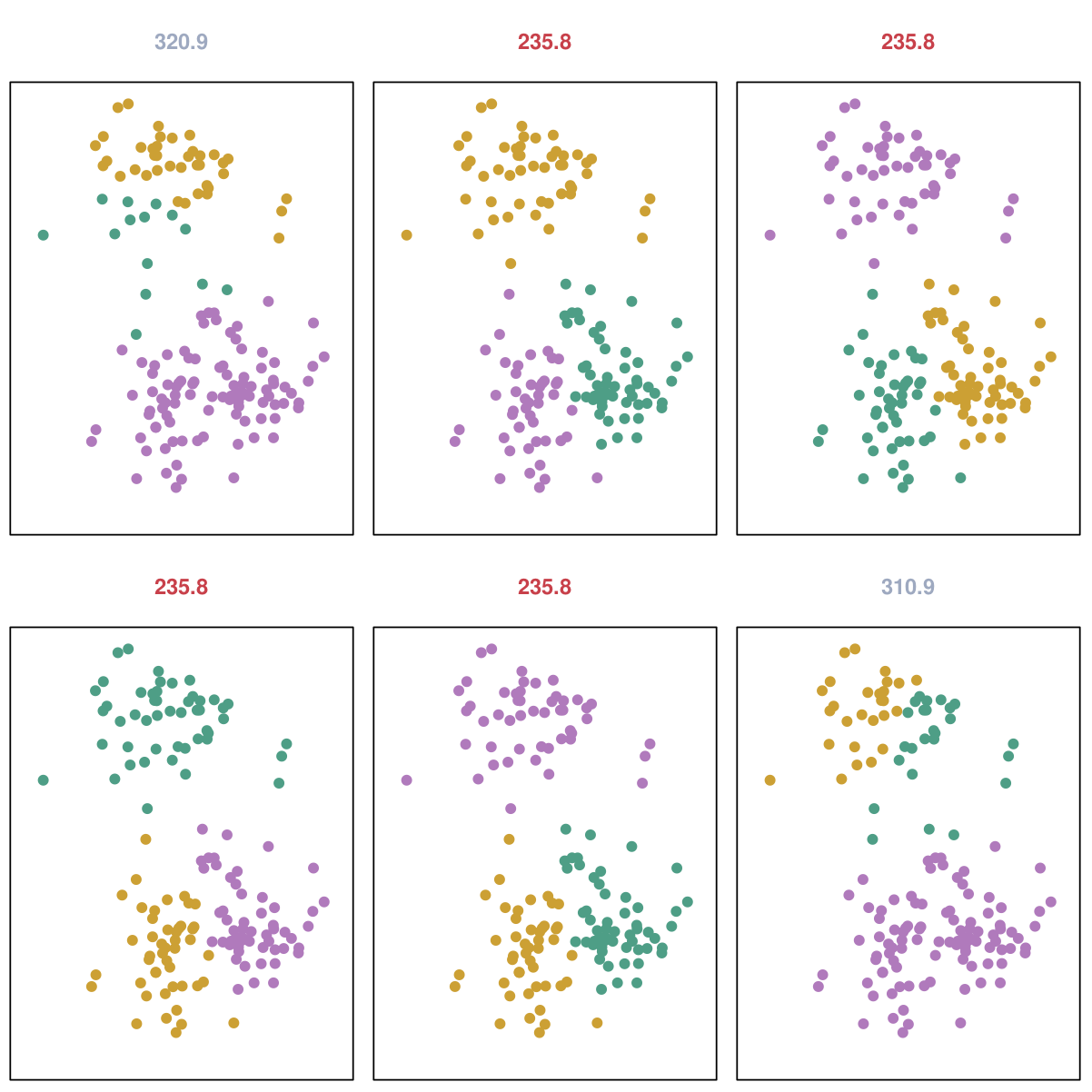
In practice, common to start from many random initializations and choose the output which minimizes the objective function.
Choosing the number of clusters: gap statistic#

Basic idea: after we have discovered all the clusters, the drop in sums of squares will be similar to drops on “null” data.
Requires a model for “null data”: \(N(0, I)\)? Unif\(([-1,1]^p)\)?
Choosing the number of clusters: gap statistic#
Rule:
with \(SD_{K+1}\) some estimate of the SD.
Extracting features from \(K\)-means#
Natural features to extract from \(K\)-means is the cluster assignment
with \(\bar{X}_k\) the cluster centers.
Caveats:#
Improvement in prediction does not necessarily imply structure in features.
Real structure in features (i.e. clusters) may not lead to improvement in prediction.
Variants#
Any MDS-type method yields points in Euclidean space
\(\implies\) a clean
Pipeline…
SpectralClustering = Pipeline([('embed', SpectralEmbedding(n_components=3)),
('clust', KMeans(n_clusters=5))])
Mixture modeling#
A soft clustering algorithm.
Within class model \(f_j(X;\theta_j), 1 \leq j \leq K\).
As it is a generative model we have a likelihood we can use for choosing \(K\) by BIC / AIC, etc.
Corresponds to hierarchical model with latent labels \(Z\):
Fit by well-known EM algorithm.
EM steps#
E-step: given current values \(\tilde{\Theta}=(\tilde{\theta}_1, \dots, \tilde{\theta}_K, \tilde{\pi}_1, \dots, \tilde{\pi}_K)\)
M-step (when no constraints on \(\theta_j\) i.e. not shared across components):
Choices to be made for mixture#
We must choose a \(K\) (BIC / AIC?)
If \(f(\cdot, \theta_j) = N(\mu_j, \Sigma_j)\) we still need to choose a model for \(\Sigma_j\) between and within classes.
Between classes: typically choice is common or class-specific covariance?
If the same, update for \(\Sigma=\Sigma_j\) is a pooled estimate of covariance with weights…
Feature extraction#
Responsibilities \(\widehat{\gamma}_j\) can replace \(K\)-means’ classification rule as features…
Relation with \(K\)-means#
If we stick with common covariance \(\sigma^2 I\) and send \(\sigma^2 \to 0\), we see
with
Model-based clustering#
Library
mclustinR.Or
GaussianMixture.predict()
Using GaussianMixture to cluster#
\(K\)-medoid clustering#
\(K\)-means / Gaussian mixture models produce centroids that “represent” each cluster as a prototype.
Both rely on Euclidean structure in order to compute centroids: they are (weighted) class averages…
\(K\)-medoid chooses instead a sample point as the prototype: given cluster \(\{X_j:j \in C\}\) and metric \(d\) the medoid is
Assignment step is the same as \(K\)-means.
Some variants of the assignment / swap step that are not exactly like \(K\) means (I believe these variants are used by
pambelow).In Euclidean case, said to be more robust because we use norms instead of squared norms.
from sklearn_extra.cluster import KMedoids
Hierarchical clustering#

Most algorithms for hierarchical clustering are bottom up.
Agglomerative process summarized by a dendrogram.
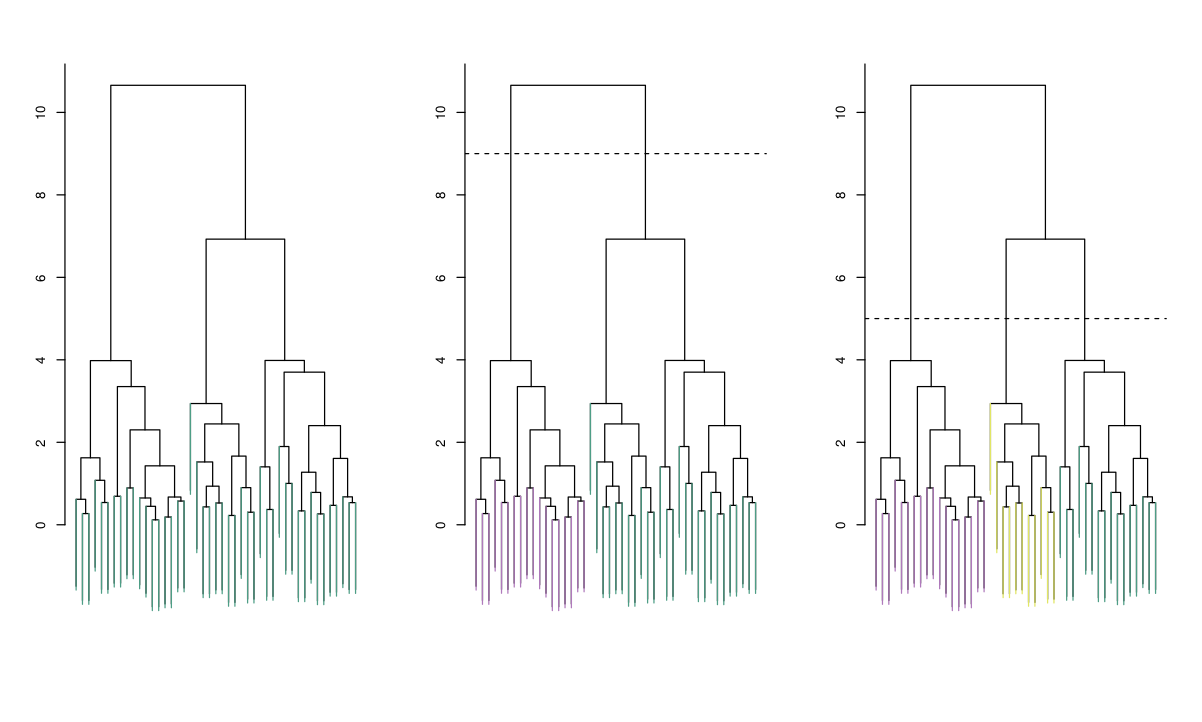
Clustering is completely determined by initial distance (or dissimilarity) matrix and the choice of dissimilarity between clusters.
The number of clusters is not fixed: each cut of the dendrogram induces a clustering.
At each step, we link the 2 clusters that are closest (least dissimilar) to each other.
Hierarchical clustering algorithms are classified according to the notion of distance between clusters.
 |
 |
 |
Complete linkage: The distance between 2 clusters is the maximum distance between any pair of samples, one in each cluster.
Single linkage: The distance between 2 clusters is the minimum distance between any pair of samples, one in each cluster.
Single linkage suffers from the chaining phenomenon.
Single linkage enjoys a continuity property the others are not known to.
Average linkage: The distance between 2 clusters is the average of all pairwise distances.
Complete linkage#
Average linkage#
Single linkage#
Choices to make#
Because it is so general, there are many choices to make in hierarchical clustering.
Choice of initial dissimilarity matrix is obviously important: e.g. do we measure distance modulo scale?
Choice of linkage also clearly makes a difference.
Some properties of hierarchical clustering#
Single, complete and average linkage have no inversions: heights when parents are merged are higher than any of their children.
Both single and complete are monotone: monotone transformations of the distances do not change structure of the trees. Not necessarily true for average linkage.
Both single and complete ignore duplicates: duplications of points do not change the structure of the true – many merges happen at height 0. Not necessarily true for average linkage.
Hierarchical clustering with prototypes: minimax linkage#
One of the nice features about \(K\)-means / \(K\)-medoids / mixture modeling is that each cluster has a simple representative: \(\implies\) useful for downstream tasks…
Bien and Tibshirani consider a different dissimilarity between clustering: minimax linkage
Based on
with the minimizer in determining \(r(C)\) the prototype for \(C\).
Dissimilarity between clusters $\( D(G,H) = r(G \cup H) \)$
This linkage is monotone, ignores density and has no inversions.

Source: Bien and Tibshirani

Source: Bien and Tibshirani