Bootstrap#
Download#
Bootstrapping linear regression#
 {height=400 fig-align=”center”}
{height=400 fig-align=”center”}
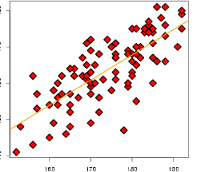
Figure: our multiple linear regression model#
We’ve talked about checking assumptions.
What to do if the assumptions don’t hold?
We will use the bootstrap!
Random \(X\): pairs bootstrap#
Suppose we think of the pairs \((X_i, Y_i)\) coming from some distribution \(F\) – this is a distribution for both the features and the outcome.
In our usual model, \(\beta\) is clearly defined. What is \(\beta\) without this assumption?
Population least squares#
For our distribution \(F\), we can define
where \((\pmb{X}, \pmb{Y}) \sim F\) leading to
Asymptotics#
In fact, our least squares estimator is \(\beta(\hat{F}_n)\) where \(\hat{F}_n\) is the empirical distribution of our sample of \(n\) observations from \(F\).
As we take a larger and larger sample,
and
for some covariance matrix \(\Sigma=\Sigma(F)\) depending only on \(F\).
Recall the variance of OLS estimator (with \(X\) fixed): $\( (X^TX)^{-1} \text{Var}(X^T(Y-X\beta)) (X^TX)^{-1}. \)$
With \(X\) random and \(n\) large this is approximately $\( \frac{1}{n} \left(E_F[\pmb{X}\pmb{X}^T] \right)^{-1} \text{Var}_F(\pmb{X} \cdot (\pmb{Y} - \pmb{X} \beta(F))) \left(E_F[\pmb{X}\pmb{X}^T] \right)^{-1}. \)$
Population least squares#
In usual model,
This is wrong in general!
We will use OLS estimate – but correct its variance!
Can we get our hands on \(\text{Var}(X^T(Y-X\beta))\) or \(\text{Var}(\hat{\beta})\) without a model?
Nonparametric bootstrap in a nutshell#
Basic algorithm for pairs#
There are many variants of the bootstrap, most using roughly this structure
boot_sample = c()
for (b in 1:B) {
idx_star = sample(1:n, n, replace=TRUE)
X_star = X[idx_star,]
Y_star = Y[idx_star]
boot_sample = rbind(boot_sample, coef(lm(Y_star ~ X_star)))
}
cov_beta_boot = cov(boot_sample)
Residual bootstrap#
If \(X\) is fixed, it doesn’t make sense to sample new \(X\) values for \(X^*\).
Residual bootstrap keeps \(X\) fixed, but adds randomly sampled residuals
boot_sample = c()
M = lm(Y ~ X - 1)
beta.hat = coef(M)
X = model.matrix(M)
Y.hat = X @ beta.hat
r.hat = Y - Y.hat
for (b in 1:B) {
idx_star = sample(1:n, n, replace=TRUE)
X_star = X
Y_star = Y.hat + r.hat[idx_star]
boot_sample = rbind(boot_sample, coef(lm(Y_star ~ X_star - 1)))
}
cov_beta_boot = cov(boot_sample)
Bootstrap for inference#
Estimated covariance
cov_beta_bootcan be used to estimate \(\text{Var}(a^T\hat{\beta})\) for confidence intervals or general linear hypothesis tests.Software does something slightly different – using percentiles of the bootstrap sample: bootstrap percentile intervals.
Bootstrapping regression#
Reference for more R examples#
library(car)
n = 50
X = rexp(n)
Y = 3 + 2.5 * X + X * (rexp(n) - 1) # our usual model is false here!
Y.lm = lm(Y ~ X)
Loading required package: carData
pairs.Y.lm = Boot(Y.lm, coef, method='case', R=1000)
confint(Y.lm)
confint(pairs.Y.lm, type='norm') # using bootstrap SE
confint(pairs.Y.lm, type='perc') # using percentiles
Loading required namespace: boot
| 2.5 % | 97.5 % | |
|---|---|---|
| (Intercept) | 2.501869 | 3.353018 |
| X | 2.235786 | 2.978521 |
| 2.5 % | 97.5 % | |
|---|---|---|
| (Intercept) | 2.626767 | 3.237522 |
| X | 2.021954 | 3.179938 |
| 2.5 % | 97.5 % | |
|---|---|---|
| (Intercept) | 2.565797 | 3.178609 |
| X | 2.135828 | 3.279040 |
Using the boot package#
The
Bootfunction incaris a wrapper around the more generalbootfunction.
library(boot)
D = data.frame(X, Y)
bootstrap_stat = function(D, bootstrap_idx) {
return(summary(lm(Y ~ X, data=D[bootstrap_idx,]))$coef[,1])
}
boot_results = boot(D, bootstrap_stat, R=500)
confint(boot_results, type='norm') # using bootstrap SE
confint(boot_results, type='perc') # using percentiles
Attaching package: ‘boot’
The following object is masked from ‘package:car’:
logit
| 2.5 % | 97.5 % |
|---|---|
| 2.613870 | 3.233349 |
| 2.021205 | 3.204503 |
| 2.5 % | 97.5 % |
|---|---|
| 2.556250 | 3.170863 |
| 2.110186 | 3.275172 |
How is the coverage?#
First we’ll use the standard regression model but errors that aren’t Gaussian.
noise = function(n) { return(rexp(n) - 1) }
simulate_correct = function(n=200, b=0.5) {
X = rexp(n)
Y = 3 + b * X + noise(n)
Y.lm = lm(Y ~ X)
# parametric interval
int_param = confint(Y.lm)[2,]
# pairs bootstrap interval
pairs.Y.lm = Boot(Y.lm, coef, method='case', R=500)
pairs_SE = sqrt(cov(pairs.Y.lm$t)[2,2]) # $t is the bootstrap sample
int_pairs = c(coef(Y.lm)[2] - qnorm(0.975) * pairs_SE,
coef(Y.lm)[2] + qnorm(0.975) * pairs_SE)
result = c((int_param[1] < b) * (int_param[2] > b),
(int_pairs[1] < b) * (int_pairs[2] > b))
names(result) = c('parametric', 'bootstrap')
return(result)
}
Check one instance#
simulate_correct()
- parametric
- 1
- bootstrap
- 1
Check coverage#
nsim = 100
coverage = c()
for (i in 1:nsim) {
coverage = rbind(coverage, simulate_correct())
}
print(apply(coverage, 2, mean))
parametric bootstrap
0.94 1.00
Misspecified model#
Now we make data for which we might have used WLS but we don’t have a model for the weights!
simulate_incorrect = function(n=200, b=0.5) {
X = rexp(n)
# the usual model is
# quite off here -- Var(X^TY) is not well
# approximated by sigma^2 * X^TX...
Y = 3 + b * X + X * noise(n)
Y.lm = lm(Y ~ X)
# parametric interval
int_param = confint(Y.lm)[2,]
# pairs bootstrap interval
pairs.Y.lm = Boot(Y.lm, coef, method='case', R=500)
pairs_SE = sqrt(cov(pairs.Y.lm$t)[2,2]) # $t is the bootstrap sample of coefficients
# we want the 2nd coefficient
int_pairs = c(coef(Y.lm)[2] - qnorm(0.975) * pairs_SE,
coef(Y.lm)[2] + qnorm(0.975) * pairs_SE)
result = c((int_param[1] < b) * (int_param[2] > b),
(int_pairs[1] < b) * (int_pairs[2] > b))
names(result) = c('parametric', 'bootstrap')
return(result)
}
Check one instance#
simulate_incorrect()
- parametric
- 0
- bootstrap
- 0
Check coverage#
nsim = 100
coverage = c()
for (i in 1:nsim) {
coverage = rbind(coverage, simulate_incorrect())
}
print(apply(coverage, 2, mean))
parametric bootstrap
0.58 0.93