Logistic regression#
Download#
Outline#
Case studies:
A. Survival in the Donner party
B. Smoking and bird-keeping
Binary outcomes
Estimation and inference: likelihood
Case study A: Donner party#
url = 'https://raw.githubusercontent.com/StanfordStatistics/stats191-data/main/Sleuth3/donner_party.csv'
donner.df = read.csv(url, sep=',', header=TRUE)
head(donner.df)
| Age | Sex | Status | |
|---|---|---|---|
| <int> | <chr> | <chr> | |
| 1 | 23 | Male | Died |
| 2 | 40 | Female | Survived |
| 3 | 40 | Male | Survived |
| 4 | 30 | Male | Died |
| 5 | 28 | Male | Died |
| 6 | 40 | Male | Died |
Binary outcomes#
Most models so far have had response \(Y\) as continuous.
StatusisDied/SurvivedOther examples:
medical: presence or absence of cancer
financial: bankrupt or solvent
industrial: passes a quality control test or not
Modelling probabilities#
For \(0-1\) responses we need to model
That is, \(Y\) is Bernoulli with a probability that depends on covariates \(\{X_1, \dots, X_p\}.\)
Note: \(\text{Var}(Y|X) = \pi(X) ( 1 - \pi(X)) = E(Y|X) \cdot ( 1- E(Y|X))\)
Or, the binary nature forces a relation between mean and variance of \(Y\).
This makes logistic regression a
Generalized Linear Model.
A possible model for Donner#
Set
Y=Status=='Survived'Simple model:
lm(Y ~ Age + Sex)Problems / issues:
We must have \(0 \leq E(Y|X) = \pi(X_1,X_2) \leq 1\). OLS will not force this.
Inference methods for will not work because of relation between mean and variance. Need to change how we compute variance!
Logistic regression#
Set
X_1=Age,X_2=SexLogistic model
This automatically fixes \(0 \leq E(Y|X) = \pi(X_1,X_2) \leq 1\).
Logit transform#
Definition: \(\text{logit}(\pi(X_1, X_2)) = \log\left(\frac{\pi(X_1, X_2)}{1 - \pi(X_1,X_2)}\right) = \beta_0 + \beta_1 X_1 + \beta_2 X_2\)
Logistic distribution#
logit.inv = function(x) {
return(exp(x) / (1 + exp(x)))
}
x = seq(-4, 4, length=200)
plot(x, logit.inv(x), lwd=2, type='l', col='red', cex.lab=1.2)
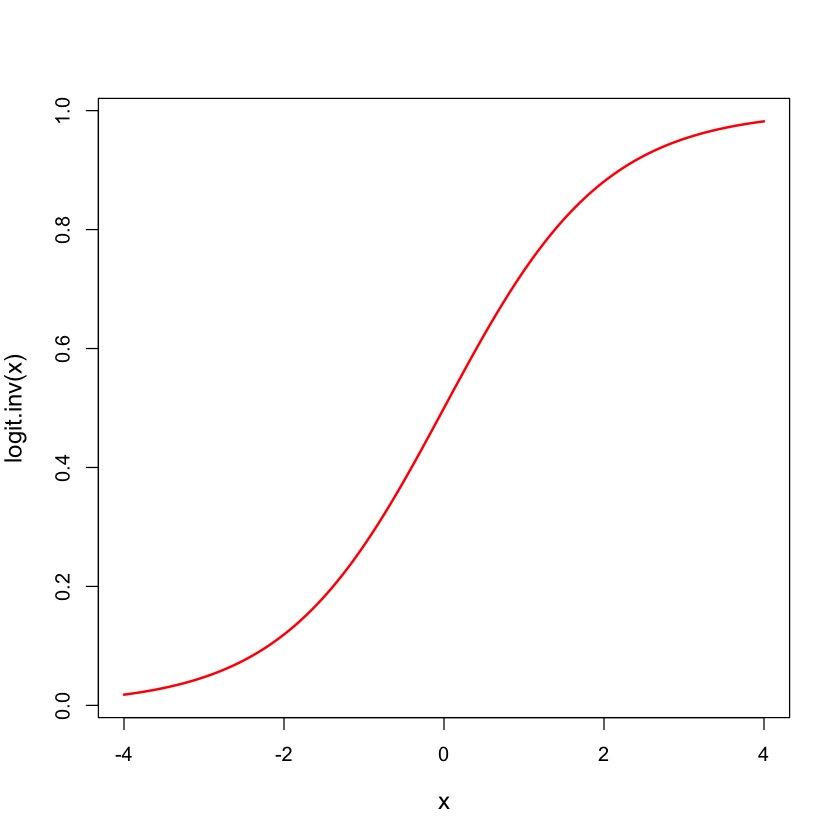
Logistic transform: logit#
logit = function(p) {
return(log(p / (1 - p)))
}
p = seq(0.01,0.99,length=200)
plot(p, logit(p), lwd=2, type='l', col='red', cex.lab=1.2)
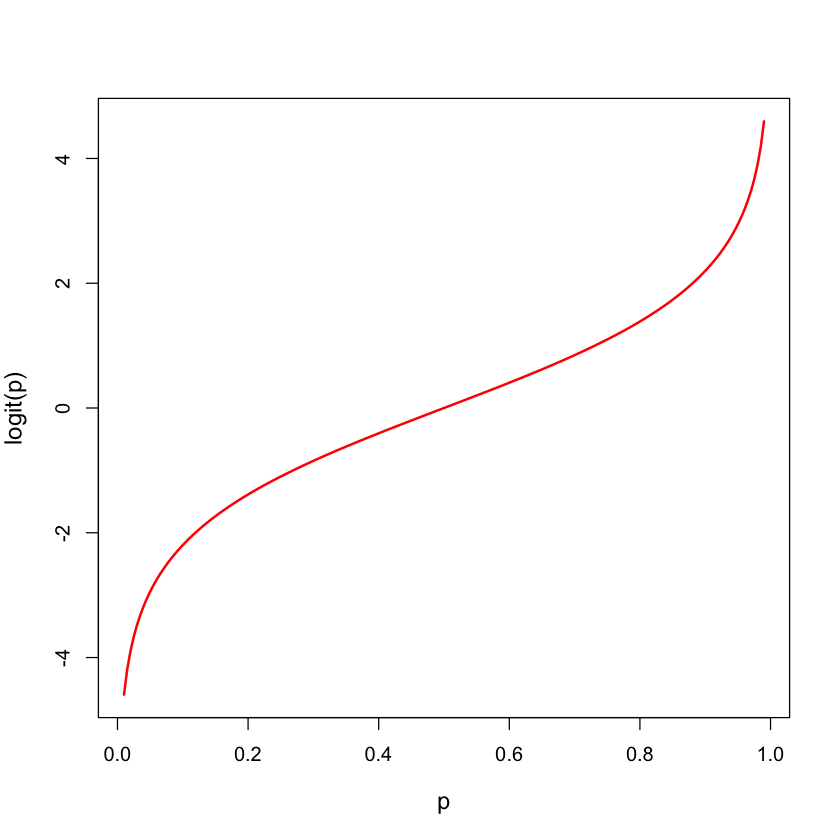
Binary regression models#
Models \(E(Y)\) as \(F(\beta_0 + \beta_1 X_1 + \beta_2 X_2)\) for some increasing function \(F\) (usually a distribution function).
The logistic model uses the function (we called
logit.invabove)
Can be fit using Maximum Likelihood / Iteratively Reweighted Least Squares.
For logistic regression, coefficients have nice interpretation in terms of
odds ratios(to be defined shortly).What about inference?
Criterion used to fit model#
Instead of sum of squares, logistic regression uses deviance:
\(DEV(\mu| Y) = -2 \log L(\mu| Y) + 2 \log L(Y| Y)\) where \(\mu\) is a location estimator for \(Y\).
If \(Y\) is Gaussian with independent \(N(\mu_i,\sigma^2)\) entries then \(DEV(\mu| Y) = \frac{1}{\sigma^2}\sum_{i=1}^n(Y_i - \mu_i)^2\)
If \(Y\) is a binary vector, with mean vector \(\pi\) then \(DEV(\pi| Y) = -2 \sum_{i=1}^n \left( Y_i \log(\pi_i) + (1-Y_i) \log(1-\pi_i) \right)\)
Minimizing deviance \(\iff\) Maximum Likelihood
Deviance for logistic regression#
For any binary regression model, \(\pi=\pi(\beta)\).
The deviance is:
For the logistic model, the RHS is:
The logistic model is special in that \(\text{logit}(\pi(\beta))=X\beta\). If we used a different transformation, the first part would not be linear in \(X\beta\).
donner.df$Y = donner.df$Status == 'Survived'
donner.glm = glm(Y ~ Age + Sex, data=donner.df,
family=binomial())
summary(donner.glm)
Call:
glm(formula = Y ~ Age + Sex, family = binomial(), data = donner.df)
Coefficients:
Estimate Std. Error z value Pr(>|z|)
(Intercept) 3.23041 1.38686 2.329 0.0198 *
Age -0.07820 0.03728 -2.097 0.0359 *
SexMale -1.59729 0.75547 -2.114 0.0345 *
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
(Dispersion parameter for binomial family taken to be 1)
Null deviance: 61.827 on 44 degrees of freedom
Residual deviance: 51.256 on 42 degrees of freedom
AIC: 57.256
Number of Fisher Scoring iterations: 4
Odds Ratios#
One reason logistic models are popular is that the parameters have simple interpretations in terms of odds
\[ODDS(A) = \frac{P(A)}{1-P(A)}.\]Logistic model:
If \(X_j \in {0, 1}\) is dichotomous, then odds for group with \(X_j = 1\) are \(e^{\beta_j}\) higher, other parameters being equal.
Tempting to think of odds ratio as probability ratio, but not quite the same
In Donner, the odds ratio for a 45 year old male compared to a 35 year old female
logodds = predict(donner.glm, list(Age=c(35,45),Sex=c('Female', 'Male')),
type='link')
logodds
logodds[2] - logodds[1]
exp(logodds[2] - logodds[1])
- 1
- 0.493271271049863
- 2
- -1.88606295597673
The estimated probabilities yield a ratio of \(0.1317/0.6209 \approx 0.21\).
prob = exp(logodds)/(1+exp(logodds))
prob
prob[2] / prob[1]
- 1
- 0.620876756856824
- 2
- 0.131694020754108
When success is rare (rare disease hypothesis) then odds ratios are approximately probability ratios…
Inference#
Model is fit to minimize deviance
Fit using IRLS (Iteratively Reweighted Least Squares) algorithm
General formula for \(\text{Var}(\hat{\beta})\) from IRLS.
Intervals formed width
vcov()are called Wald intervals.
center = coef(donner.glm)['Age']
SE = sqrt(vcov(donner.glm)['Age', 'Age'])
U = center + SE * qnorm(0.975)
L = center - SE * qnorm(0.975)
data.frame(L, center, U)
| L | center | U | |
|---|---|---|---|
| <dbl> | <dbl> | <dbl> | |
| Age | -0.1512803 | -0.07820407 | -0.005127799 |
Covariance#
The estimated covariance uses the “weights” computed from the fitted model.
pi.hat = fitted(donner.glm)
W.hat = pi.hat * (1 - pi.hat)
X = model.matrix(donner.glm)
C = solve(t(X) %*% (W.hat * X))
c(SE, sqrt(C['Age', 'Age']))
- 0.0372844979677129
- 0.0372874889556373
Confidence intervals#
The intervals above are slightly different from what
Rwill give you if you ask it for confidence intervals.Ruses so-called profile intervals.For large samples the two methods should agree quite closely.
CI = confint(donner.glm)
CI
mean(CI[2,]) # profile intervals are not symmetric around the estimate...
data.frame(L, center, U)
Waiting for profiling to be done...
| 2.5 % | 97.5 % | |
|---|---|---|
| (Intercept) | 0.8514190 | 6.42669512 |
| Age | -0.1624377 | -0.01406576 |
| SexMale | -3.2286705 | -0.19510198 |
| L | center | U | |
|---|---|---|---|
| <dbl> | <dbl> | <dbl> | |
| Age | -0.1512803 | -0.07820407 | -0.005127799 |
Example from book#
The book considers testing whether there is any interaction between
SexandAgein the Donner party data:
summary(glm(Y ~ Age + Sex + Age:Sex,
donner.df,
family=binomial()))$coef
| Estimate | Std. Error | z value | Pr(>|z|) | |
|---|---|---|---|---|
| (Intercept) | 7.2463848 | 3.20516725 | 2.260845 | 0.02376889 |
| Age | -0.1940741 | 0.08741539 | -2.220137 | 0.02640948 |
| SexMale | -6.9280495 | 3.39887166 | -2.038338 | 0.04151614 |
| Age:SexMale | 0.1615969 | 0.09426191 | 1.714339 | 0.08646650 |
\(Z\)-test for testing \(H_0:\beta_{\texttt Age:SexMale}=0\):
When \(H_0\) is true, \(Z\) is approximately \(N(0,1)\).
pval = 2 * pnorm(1.714, lower=FALSE)
pval
Comparing models in logistic regression#
For a model \({\cal M}\), \(DEV({\cal M})\) replaces \(SSE({\cal M})\).
In least squares regression (with \(\sigma^2\) known), we use
This is closely related to \(F\) with large \(df_F\): approximately \(F_{df_R-df_F, df_F} \cdot (df_R-df_F)\).
For logistic regression this difference in \(SSE\) is replaced with
Resulting tests do not agree numerically with those coming from IRLS (Wald tests). Both are often used.
Comparing ~ Age + Sex to ~1#
anova(glm(Y ~ 1,
data=donner.df,
family=binomial()),
donner.glm)
| Resid. Df | Resid. Dev | Df | Deviance | Pr(>Chi) | |
|---|---|---|---|---|---|
| <dbl> | <dbl> | <dbl> | <dbl> | <dbl> | |
| 1 | 44 | 61.82654 | NA | NA | NA |
| 2 | 42 | 51.25628 | 2 | 10.57026 | 0.00506638 |
Corresponding \(p\)-value:
1 - pchisq(10.57, 2) # testing ~1 vs ~1 + Health.Aware + Age
Comparing ~ Age + Sex to ~ Age#
anova(glm(Y ~ Age,
data=donner.df,
family=binomial()),
donner.glm)
| Resid. Df | Resid. Dev | Df | Deviance | Pr(>Chi) | |
|---|---|---|---|---|---|
| <dbl> | <dbl> | <dbl> | <dbl> | <dbl> | |
| 1 | 43 | 56.29072 | NA | NA | NA |
| 2 | 42 | 51.25628 | 1 | 5.034437 | 0.02484816 |
Corresponding \(p\)-value:
1 - pchisq(5.03, 1) # testing ~ 1 + Health.Aware vs ~1 + Health.Aware + Age
Let’s compare this with the Wald test:
summary(donner.glm)
Call:
glm(formula = Y ~ Age + Sex, family = binomial(), data = donner.df)
Coefficients:
Estimate Std. Error z value Pr(>|z|)
(Intercept) 3.23041 1.38686 2.329 0.0198 *
Age -0.07820 0.03728 -2.097 0.0359 *
SexMale -1.59729 0.75547 -2.114 0.0345 *
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
(Dispersion parameter for binomial family taken to be 1)
Null deviance: 61.827 on 44 degrees of freedom
Residual deviance: 51.256 on 42 degrees of freedom
AIC: 57.256
Number of Fisher Scoring iterations: 4
Diagnostics#
Similar to least square regression, only residuals used are usually deviance residuals
\[r_i = \text{sign}(Y_i-\widehat{\pi}_i) \sqrt{DEV(\widehat{\pi}_i|Y_i)}.\]These agree with usual residual for least square regression.
par(mfrow=c(2,2))
plot(donner.glm)

influence.measures(donner.glm)
Influence measures of
glm(formula = Y ~ Age + Sex, family = binomial(), data = donner.df) :
dfb.1_ dfb.Age dfb.SxMl dffit cov.r cook.d hat inf
1 -0.0828 0.09169 -0.0900 -0.2341 1.047 0.015372 0.0492
2 0.0228 0.13667 -0.2285 0.3671 1.100 0.038310 0.1025
3 -0.2909 0.32202 0.2497 0.4314 0.916 0.094717 0.0567
4 0.0308 -0.03411 -0.0974 -0.1648 1.065 0.006840 0.0387
5 0.0047 -0.00520 -0.0971 -0.1763 1.058 0.008042 0.0389
6 0.0972 -0.10766 -0.0835 -0.1442 1.111 0.004763 0.0567
7 0.0701 -0.23190 0.1880 -0.3990 1.146 0.043320 0.1310
8 0.0599 -0.06636 -0.0309 -0.0697 1.146 0.001030 0.0670
9 0.0518 -0.05738 -0.0259 -0.0598 1.143 0.000756 0.0628
10 0.0701 -0.23190 0.1880 -0.3990 1.146 0.043320 0.1310
11 -0.3939 0.23169 0.4066 -0.4886 0.955 0.109490 0.0780
12 -0.0071 0.00786 0.1466 0.2663 0.986 0.024482 0.0389
13 0.0047 -0.00520 -0.0971 -0.1763 1.058 0.008042 0.0389
14 -0.0828 0.09169 -0.0900 -0.2341 1.047 0.015372 0.0492
15 0.1534 -0.10271 -0.1414 0.1748 1.139 0.006983 0.0802
16 0.1543 -0.09959 -0.1473 0.1801 1.136 0.007465 0.0794
17 -0.0071 0.00786 0.1466 0.2663 0.986 0.024482 0.0389
18 0.1342 -0.10552 -0.1022 0.1391 1.155 0.004259 0.0839
19 0.1005 -0.25632 0.1656 -0.3986 1.172 0.041875 0.1438
20 0.0744 -0.08240 -0.0408 -0.0880 1.150 0.001656 0.0721
21 0.1500 -0.10660 -0.1298 0.1644 1.145 0.006103 0.0817
22 0.1714 -0.18980 0.0614 0.2872 1.076 0.022793 0.0735
23 -0.0439 0.04860 -0.0940 -0.2054 1.050 0.011427 0.0433
24 0.0656 -0.07267 -0.0346 -0.0768 1.148 0.001254 0.0694
25 0.0557 -0.06172 0.1193 0.2608 1.009 0.021757 0.0433
26 0.1434 -0.15882 0.0768 0.2773 1.055 0.022058 0.0623
27 -0.0992 0.10982 0.1833 0.2998 0.962 0.034818 0.0403
28 0.1259 -0.02425 -0.1980 0.2436 1.109 0.014897 0.0786
29 0.1546 -0.09559 -0.1532 0.1855 1.133 0.007983 0.0787
30 -0.0522 0.05781 0.1650 0.2793 0.974 0.028470 0.0387
31 -0.2884 0.31929 -0.0567 -0.4225 1.046 0.059528 0.0928
32 0.1370 -0.28212 0.1339 -0.3937 1.211 0.039152 0.1626
33 0.1520 -0.10503 -0.1355 0.1695 1.142 0.006530 0.0810
34 -0.0439 0.04860 -0.0940 -0.2054 1.050 0.011427 0.0433
35 -0.4172 0.46184 0.2851 0.5385 0.876 0.187509 0.0685
36 0.1259 -0.02425 -0.1980 0.2436 1.109 0.014897 0.0786
37 0.0308 -0.03411 -0.0974 -0.1648 1.065 0.006840 0.0387
38 -0.0439 0.04860 -0.0940 -0.2054 1.050 0.011427 0.0433
39 -0.0439 0.04860 -0.0940 -0.2054 1.050 0.011427 0.0433
40 -0.0439 0.04860 -0.0940 -0.2054 1.050 0.011427 0.0433
41 0.0308 -0.03411 -0.0974 -0.1648 1.065 0.006840 0.0387
42 0.0761 -0.08422 -0.0931 -0.1518 1.087 0.005495 0.0453
43 0.0937 -0.10368 0.1018 0.2648 1.026 0.021407 0.0492
44 -0.0627 0.06942 -0.0922 -0.2188 1.049 0.013186 0.0459
45 0.1541 -0.09064 -0.1591 0.1912 1.130 0.008544 0.0780
Model selection#
Models all have AIC scores \(\implies\) stepwise selection can be used easily!
Package
bestglmhas something likeleapsfunctionality (not covered here)
AIC(donner.glm)
null.glm = glm(Y ~ 1, data=donner.df, family=binomial())
upper.glm = glm(Y ~ (.-Status)^2, data=donner.df, family=binomial())
donner_selected = step(null.glm, scope=list(upper=upper.glm), direction='both')
summary(donner_selected)$coef
Start: AIC=63.83
Y ~ 1
Df Deviance AIC
+ Age 1 56.291 60.291
+ Sex 1 57.286 61.286
<none> 61.827 63.827
Step: AIC=60.29
Y ~ Age
Df Deviance AIC
+ Sex 1 51.256 57.256
<none> 56.291 60.291
- Age 1 61.827 63.827
Step: AIC=57.26
Y ~ Age + Sex
Df Deviance AIC
+ Age:Sex 1 47.346 55.346
<none> 51.256 57.256
- Sex 1 56.291 60.291
- Age 1 57.286 61.286
Step: AIC=55.35
Y ~ Age + Sex + Age:Sex
Df Deviance AIC
<none> 47.346 55.346
- Age:Sex 1 51.256 57.256
| Estimate | Std. Error | z value | Pr(>|z|) | |
|---|---|---|---|---|
| (Intercept) | 7.2463848 | 3.20516725 | 2.260845 | 0.02376889 |
| Age | -0.1940741 | 0.08741539 | -2.220137 | 0.02640948 |
| SexMale | -6.9280495 | 3.39887166 | -2.038338 | 0.04151614 |
| Age:SexMale | 0.1615969 | 0.09426191 | 1.714339 | 0.08646650 |
Case Study B: birdkeeping and smoking#
Outcome:
LC, lung cancer statusFeatures:
FM: sex of patientAG: age (years)SS: socio-economic statusYR: years of smoking prior to examination / diagnosisCD: cigarettes per dayBK: whether or not subjects keep birds at home
url = 'https://raw.githubusercontent.com/StanfordStatistics/stats191-data/main/Sleuth3/smoking_birds.csv'
smoking.df = read.csv(url, sep=',', header=TRUE)
smoking.df$LC = smoking.df$LC == 'LungCancer'
head(smoking.df)
| LC | FM | SS | BK | AG | YR | CD | |
|---|---|---|---|---|---|---|---|
| <lgl> | <chr> | <chr> | <chr> | <int> | <int> | <int> | |
| 1 | TRUE | Male | Low | Bird | 37 | 19 | 12 |
| 2 | TRUE | Male | Low | Bird | 41 | 22 | 15 |
| 3 | TRUE | Male | High | NoBird | 43 | 19 | 15 |
| 4 | TRUE | Male | Low | Bird | 46 | 24 | 15 |
| 5 | TRUE | Male | Low | Bird | 49 | 31 | 20 |
| 6 | TRUE | Male | High | NoBird | 51 | 24 | 15 |
base.glm = glm(LC ~ .,
data=smoking.df,
family=binomial())
summary(base.glm)$coef
| Estimate | Std. Error | z value | Pr(>|z|) | |
|---|---|---|---|---|
| (Intercept) | 0.09195626 | 1.75464706 | 0.05240727 | 0.9582041827 |
| FMMale | -0.56127270 | 0.53116056 | -1.05669122 | 0.2906525319 |
| SSLow | -0.10544761 | 0.46884614 | -0.22490877 | 0.8220502474 |
| BKNoBird | -1.36259456 | 0.41127585 | -3.31309159 | 0.0009227076 |
| AG | -0.03975542 | 0.03548022 | -1.12049525 | 0.2625027758 |
| YR | 0.07286848 | 0.02648741 | 2.75106120 | 0.0059402544 |
| CD | 0.02601689 | 0.02552400 | 1.01931102 | 0.3080553359 |
Reduced model: ~ . - BK#
reduced.glm = glm(LC ~ . - BK,
data=smoking.df,
family=binomial())
anova(reduced.glm, base.glm)
| Resid. Df | Resid. Dev | Df | Deviance | Pr(>Chi) | |
|---|---|---|---|---|---|
| <dbl> | <dbl> | <dbl> | <dbl> | <dbl> | |
| 1 | 141 | 165.8684 | NA | NA | NA |
| 2 | 140 | 154.1984 | 1 | 11.66997 | 0.000635172 |
Further reduction: ~ FM + SS + AG + YR#
reduced2.glm = glm(LC ~ . - BK - CD,
data=smoking.df,
family=binomial())
anova(reduced2.glm, reduced.glm)
| Resid. Df | Resid. Dev | Df | Deviance | Pr(>Chi) | |
|---|---|---|---|---|---|
| <dbl> | <dbl> | <dbl> | <dbl> | <dbl> | |
| 1 | 142 | 166.5320 | NA | NA | NA |
| 2 | 141 | 165.8684 | 1 | 0.6636412 | 0.4152774 |
Effect of smoking#
As in the salary discrimination study, the book first selects model without
BKWe’ll start with all possible two-way interactions
full.glm = glm(LC ~ (. - BK)^2, data=smoking.df,
family=binomial())
summary(full.glm)$coef
| Estimate | Std. Error | z value | Pr(>|z|) | |
|---|---|---|---|---|
| (Intercept) | -25.905104866 | 8.969226e+02 | -0.02888221 | 0.9769585 |
| FMMale | 19.536848849 | 8.968999e+02 | 0.02178264 | 0.9826213 |
| SSLow | 18.217486968 | 8.969003e+02 | 0.02031161 | 0.9837948 |
| AG | 0.149767554 | 1.434516e-01 | 1.04402869 | 0.2964721 |
| YR | 0.261575427 | 2.351992e-01 | 1.11214402 | 0.2660762 |
| CD | 0.085858326 | 2.404498e-01 | 0.35707387 | 0.7210365 |
| FMMale:SSLow | -15.808049058 | 8.968883e+02 | -0.01762544 | 0.9859377 |
| FMMale:AG | -0.131426404 | 9.613134e-02 | -1.36715462 | 0.1715768 |
| FMMale:YR | 0.144263967 | 9.406420e-02 | 1.53367556 | 0.1251095 |
| FMMale:CD | -0.102850561 | 9.575896e-02 | -1.07405683 | 0.2827972 |
| SSLow:AG | -0.058669167 | 9.501203e-02 | -0.61749200 | 0.5369103 |
| SSLow:YR | 0.068748587 | 7.340861e-02 | 0.93651945 | 0.3490058 |
| SSLow:CD | -0.084255122 | 5.797420e-02 | -1.45332091 | 0.1461347 |
| AG:YR | -0.004907733 | 3.730934e-03 | -1.31541662 | 0.1883699 |
| AG:CD | 0.002626962 | 4.682346e-03 | 0.56103551 | 0.5747733 |
| YR:CD | -0.002597374 | 3.145647e-03 | -0.82570428 | 0.4089719 |
select.glm = step(full.glm,
direction='backward',
trace=FALSE)
# these effects aren't to be fully trusted - data snooping!
summary(select.glm)$coef
| Estimate | Std. Error | z value | Pr(>|z|) | |
|---|---|---|---|---|
| (Intercept) | 0.82886135 | 1.52661870 | 0.5429393 | 0.5871715684 |
| FMMale | -0.73637859 | 0.48914336 | -1.5054453 | 0.1322096220 |
| AG | -0.06362753 | 0.03359029 | -1.8942241 | 0.0581952757 |
| YR | 0.08776499 | 0.02451891 | 3.5794810 | 0.0003442773 |
Let’s see the effect of
BKwhen added to our selected model
bird.glm = glm(LC ~ FM + AG + YR + BK, family=binomial(),
data=smoking.df)
# these effects aren't to be fully trusted - data snooping!
anova(select.glm, bird.glm)
| Resid. Df | Resid. Dev | Df | Deviance | Pr(>Chi) | |
|---|---|---|---|---|---|
| <dbl> | <dbl> | <dbl> | <dbl> | <dbl> | |
| 1 | 143 | 166.5488 | NA | NA | NA |
| 2 | 142 | 155.3212 | 1 | 11.22754 | 0.0008059231 |
Probit model#
We used \(F(x)=e^x/(1+e^x)\). There are other choices…
Probit regression model:
Above, \(\Phi\) is CDF of \(N(0,1)\), i.e. \(\Phi(t) = {\tt pnorm(t)}\), \(\Phi^{-1}(q) = {\tt qnorm}(q)\).
Regression function:
In logit, probit and cloglog \({\text{Var}}(Y|X)=\pi(X)(1-\pi(X))\) but the model for the mean is different.
Coefficients no longer have an odds ratio interpretation.
summary(glm(Y ~ Age + Sex,
data=donner.df,
family=binomial(link='probit')))
Call:
glm(formula = Y ~ Age + Sex, family = binomial(link = "probit"),
data = donner.df)
Coefficients:
Estimate Std. Error z value Pr(>|z|)
(Intercept) 1.91730 0.76438 2.508 0.0121 *
Age -0.04571 0.02076 -2.202 0.0277 *
SexMale -0.95828 0.43983 -2.179 0.0293 *
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
(Dispersion parameter for binomial family taken to be 1)
Null deviance: 61.827 on 44 degrees of freedom
Residual deviance: 51.283 on 42 degrees of freedom
AIC: 57.283
Number of Fisher Scoring iterations: 5
Generalized linear models: glm#
Given a dataset \((Y_i, X_{i1}, \dots, X_{ip}), 1 \leq i \leq n\) we consider a model for the distribution of \(Y|X_1, \dots, X_p\).
-If \(\eta_i=g(E(Y_i|X_i)) = g(\mu_i) = \beta_0 + \sum_{j=1}^p \beta_j X_{ij}\) then \(g\) is called the link function for the model.
If \({\text{Var}}(Y_i) = \phi \cdot V({\mathbb{E}}(Y_i)) = \phi \cdot V(\mu_i)\) for \(\phi > 0\) and some function \(V\), then \(V\) is the called variance function for the model.
Canonical reference Generalized linear models.
Binary regression as GLM#
For a logistic model, \(g(\mu)={\text{logit}}(\mu), \qquad V(\mu)=\mu(1-\mu).\)
For a probit model, \(g(\mu)=\Phi^{-1}(\mu), \qquad V(\mu)=\mu(1-\mu).\)
For a cloglog model, \(g(\mu)=-\log(-\log(\mu)), \qquad V(\mu)=\mu(1-\mu).\)
All of these have dispersion \(\phi=1\).